Nature ( 2026 ) Cite this article
Although the green revolution adapted a handful of crops to homogeneous and high-input industrialized agriculture, much of the global population still relies on the local production of variable crop cultivars by low-input smallholder farms.
This diversity of unhomogenized crops 1 , like that of the grain and bioenergy crop sorghum 2 , 3 , 4 , 5 , offers raw materials for genetic gain and cultivar improvement.
However, breeding efforts can be constrained by highly specialized traits and breeding targets 6 .
Here, to bridge this diversity, we constructed a 33-member pangenome reference and a diversity panel across 1,984 cultivars and landraces.
We leveraged these resources to explore the complex interplay among historical contingency, ongoing adaptation and previously uncharacterized structural diversity.
Specifically, our analyses conclusively demonstrated multiple nested and deeply diverged structural variants in the domestication gene SHATTERING1 , which distinguish the previously established multicentric origin of sorghum.
We then applied landscape genomics to reveal how gene flow and secondary contact created the complex genetic mosaic in contemporary breeding networks.
As proof of concept for pangenome-accelerated trait discovery, we connected biosynthetic gene cluster structural variation to phenotypic leaf concentration of the cyanogenic glucoside dhurrin.
Combined, these approaches will accelerate breeding and trait discovery and provide a framework for similar applications in other crops.
Similar content being viewed by others
Extensive variation within the pan-genome of cultivated and wild sorghum
From the genome to super-pangenome: a new paradigm for accelerated crop improvement
Graph-based pangenomes and pan-phenome provide a cornerstone for eggplant biology and breeding
Sorghum ( Sorghum bicolor L.
Moench) is one of the most climate-resilient and phenotypically diverse major crops; it is adapted not only to environmental stresses but also to variable agronomic practices and end-uses 1 , 5 .
Commercial sorghum hybrids cultivated for the temperate regions of the Americas, Asia and Australia are typically single-purpose grain, forage or energy types 2 .
However, open-pollinated or inbred varieties grown by small-scale producers in tropical Africa, Asia and the Americas are typically multipurpose (grain, forage and biomass).Moreover, temperate grain sorghums were bred for reduced photoperiod sensitivity 7 , whereas small-scale producers in the tropics grow varieties that are highly photoperiod sensitive and adapted to narrow latitudinal bands 8 .
This diversity in and across sorghum gene pools can hinder modern breeding approaches, which rely on large homogeneous pools of elite breeding germplasm 9 to develop broadly adapted improved varieties.
For example, the introduction of broadly beneficial alleles into local breeding programs may alter grain characteristics or otherwise reduce the local value of a cultivar 10 .
As an example, green revolution-style sorghum varieties, developed for North America and South Asia, were brought directly into smallholder-oriented breeding programs in Africa 11 .
However, the ‘improved’ germplasm did not conform to local demands, which limited their adoption 10 .
In response to these setbacks, localized participatory breeding approaches, which engage farmers and consumers, have successfully developed locally improved sorghum varieties 3 , 4 .
The BTx623 cultivar (PI564163) has served as the reference genotype for sorghum genetics since 2009 (ref.
16 ), and its 2013 V3 genome assembly 17 remains the most important global reference resource for research and breeding.
BTx623 is also well known as a parent line for commercial grain and bioenergy hybrids in the United States 18 , the first commercial hybrids in Sudan and Niger in Central and West Africa 13 and as the genetic background for many trait discovery and pre-breeding resources 19 .
Like genome assemblies that used similar technology, the BTx623 V3 assembly contained many large gaps with unknown sequences.
Long-read genome sequencing technologies enabled us to complete the repetitive regions that typically cause such gaps.
Consequently, our updated BTx623 V5 genome assembly represents the 10 sorghum chromosomes with only 34 contigs (contig N 50 = 50.74 Mb), a 140-fold improvement over the 4,783 contigs used to assemble V3 (Supplementary Fig.
1 ; see Supplementary Note 1 for methods of the unreleased V4 assembly).
Similar to many recent upgrades of older genome sequences, repetitive regions in V5 are more extensive (for example, 2.86 times more contiguous centromere 20 blocks), whereas the gene-rich portions are moderately improved (BUSCO genome assembly completeness scores of 98.3% for V3 and 99.7% for V5).
Notably, V5 corrects several structural scaffolding errors in V3.
Although many of these large misoriented regions of V3 are highly repetitive, V5 clarifies the positions of several key genes that underlie local adaptation among breeding gene pools (Supplementary Data 1 and 2 ), including the flowering time locus Maturity1 (ref.
7 ) and the biosynthesis gene POR 21 for the secondary metabolite dhurrin.
Linkage mapping and inference of candidate genes in these regions will be more accurate with a reference that is collinear with the recombination landscape.
Reference and diversity re-sequencing panels
As an admixed breeding line, BTx623 serves as a solid reference to anchor genome-wide genetic variation, including across our diversity panel of 1,984 unique genotypes (total n = 2,145) re-sequenced with high coverage whole-genome short-read libraries (Extended Data Fig.
1 ).
This diversity panel spans substantial phenotypic variation (Fig.
1 ) for growth rate, biomass accumulation, flowering time and stress responses (Extended Data Fig.
2 ).
It also represents many existing (for example, biomass association panel (BAP), n = 375) and new (for example, the Transportation Energy Resources from Renewable Agriculture (TERRA) panel, n = 220) populations of interest, a host of germplasm groupings (breeding, n = 500) and traditional local landrace varieties ( n = 737) and 746 georeferenced genotypes with local collection coordinates (Supplementary Data 3 ).
The diversity panel also contains members of all five of the major botanical types ( n = 807 botanical assignments), which have divergent morphological traits connected to local grower preferences.
Fig.
1: Geographical and genetic distribution of the sorghum diversity panel.
The whole-genome re-sequencing panel of 1,984 unique genotypes was clustered into 10 subpopulations (Supplementary Note 4 ).
Ancestry proportions to each subpopulation are shown for all genotypes (vertical bars in the bar plot; bottom) and for the 693 unique African and Asian cultivars with georeferenced collection locations (pie charts and map; top).
The 33 members of the pangenome reference are marked with an asterisk above the bar plot.
Ten pangenome members that span the genetic and phenotypic diversity of our pangenome are labelled (including the country of origin and botanical type), with representative photographs (above the label) and their ancestry proportions (below the label).
Full size image
The whole-genome re-sequencing panel of 1,984 unique genotypes was clustered into 10 subpopulations (Supplementary Note 4 ).
Ancestry proportions to each subpopulation are shown for all genotypes (vertical bars in the bar plot; bottom) and for the 693 unique African and Asian cultivars with georeferenced collection locations (pie charts and map; top).
The 33 members of the pangenome reference are marked with an asterisk above the bar plot.
Ten pangenome members that span the genetic and phenotypic diversity of our pangenome are labelled (including the country of origin and botanical type), with representative photographs (above the label) and their ancestry proportions (below the label).
Although the BTx623 genome forms the foundation for single-reference genomic analyses, the substantial diversity across cultivated sorghum necessitates a more comprehensive approach to genomic-enabled breeding.
The recent development of multiple separately assembled genomes in many species, including sorghum 22 , has revealed functionally important large-scale sequence variation 23 that is often not perfectly captured by short reads mapped to a single reference genome 24 .
Multiple genomes, when integrated into a single pangenome reference, can be used to more accurately detect and annotate putative functional single-nucleotide polymorphism (SNP) or small insertion–deletion (indel) variants and are often necessary to confirm large-scale structural (SVs), copy-number (CNVs) and presence–absence variants (PAVs).
This aspect is especially true in sorghum and other phenotypically diverse crops with multiple deeply diverged gene pools 25 .
To facilitate these investigations, we generated a 33-member pangenome reference (Supplementary Tables 1 and 2 and see Supplementary Note 2 for detailed methods) that spans four genetic model genotypes: BTx623 V5; readily-transformable RTx430 (refs.
26 , 27 ); stay-green and drought tolerant BTx642 (ref.
26 ); and sweet Wray.
We also include 9 genotypes that span the variety of our sorghum genetic diversity panel (Fig.
1 and Extended Data Figs.
1 and 2 ) and 20 of the most important cultivars for the global sorghum improvement community.
Example cultivars include Macia and IRAT204 for breeding, CSM-63 and Mota Maradi for local adaptation, SC 35 as a stay-green line, SC 283 for tolerance to aluminium and SRN39 for resistance to the parasitic plant Striga hermonthica (hereafter referred to as striga).
We characterized phenotypic variation across the 33 reference genomes and a subset of the diversity panel by pairing standard field and physiological traits with georeferenced origin, botanical type and breeding status characteristics (Supplementary Data 4 ).
The following field and physiological traits were analysed: (1) field-collected biomass, flowering time and plant height; (2) unmanned aerial vehicle (UAV)-collected normalized difference vegetation index (NDVI) and modified chlorophyll absorption in reflectance index (MCARI); and (3) greenhouse and image-based morphology and growth, gas exchange and water use efficiency (WUE).
Combined, the reference genotypes encompass the diversity of major breeding targets such as field-based biomass accumulation and flowering time.
We also noted highly diverse phenotypes across environments and conditions (Extended Data Fig.
2 ).
For example, the lines DJOFELA and CSM-63 (cultivated and elite African breeding lines, respectively) consistently ranked highly in WUE, plant area (cm 2 ) and height (cm) under both well-watered and water-limited conditions, which highlights their potential as broadly adaptable breeding candidates.
Moreover, SRN39, another elite African line from Sudan, stood out for its strong performance under water-limited conditions, which indicated valuable drought-resilience traits.
Together, this suite of phenotypes establishes a baseline for relating selection and adaptation across landscapes to plant performance.
Global pangenome sequence and gene content
Combined, the 33-member pangenome reference nearly doubles the total sequence relative to the BTx623 linear reference and includes 325 large SVs (179.6 Mb > 100 kb) and 26 >1 Mb inversions found in multiple assemblies (Fig.
2a,b and Supplementary Data 5 ).
Outside these inversions, our sorghum genomes are collinear and lack interchromosomal translocations (for example, maize 28 ), centromeric SVs (for example, pennycress 29 ) or other factors that are common in plants and complicate pangenomic integration and breeding.
Fig.
2: Pangenome synteny map and PAV.
a , Macrosynteny map.
Syntenic blocks were calculated from windowed alignments and converted to a riparian diagram.
To aid visualization, all genomes were scaled to the same x axis extent; genome sizes are provided in Supplementary Table 1 .
b , c , Panacus ( b ) and OrthoFinder ( c ) (phylogenetically hierarchical orthogroups (OGs)) pangenome expansion curves.
In both plots, core (100% presence) and PAV elements are represented by black and grey bars, respectively.
Source data Full size image
a , Macrosynteny map.
Syntenic blocks were calculated from windowed alignments and converted to a riparian diagram.
To aid visualization, all genomes were scaled to the same x axis extent; genome sizes are provided in Supplementary Table 1 .
b , c , Panacus ( b ) and OrthoFinder ( c ) (phylogenetically hierarchical orthogroups (OGs)) pangenome expansion curves.
In both plots, core (100% presence) and PAV elements are represented by black and grey bars, respectively.
The pangenome reference also provides an opportunity to better define PAV and CNV among protein-coding genes.
However, methodological considerations are crucial for interpretation 30 .
For example, pangenomes built without independent RNA sequencing support for each genome can underestimate PAV in genes owing to a reliance on sequence similarity from related species.
Conversely, differing methods, sequencing support or statistical thresholding can inflate estimates of PAV genes through an abundance of spurious false-positive gene models 31 .
Here we sought to leverage the strengths of both approaches by first annotating all 33 members of the pangenome with deep RNA sequencing across 685 biological samples (Supplementary Data 6 ) and then removing rare gene models without strong support across any lines of evidence.
These efforts produced a pangene set with 988,756 gene models across 54,959 phylogenetically hierarchical orthogroups (gene families; Supplementary Data 7 ).
Most genes fell into the ‘core’ (100% presence; 691,667 or 70.0% of genes) or near-core (>90% presence; 101,316 or 10.2% of genes) PAV categories (Fig.
2c ).
However, some of the most important genes for domestication and ongoing crop improvement displayed PAV or CNV across the pangenome references, including the striga-resistance gene Low germination stimulant 1 ( LGS1 ; sulfotransferase, Sobic.005G213600) 32 , 33 .
Combined, variable gene content and the abundance of large-scale structural variation indicate that our representation of the sorghum pangenome may facilitate the discovery and implementation of otherwise unknown large-effect alleles.
Although pangenomes can clearly facilitate trait discovery among reference genotypes, the next step in identifying and reconstructing alleles across re-sequenced populations is both conceptually challenging and computationally intensive.
Current state-of-the-art methods require reconstruction of a pangenome graph and subsequent re-alignment of all sequencing reads back to the updated graph whenever new references are included.
Despite this burdensome computational overhead, graph-based genotyping effectively captures structural variation and reduces false variant calls (for example, pseudo-heterozygosity; Supplementary Note 3 and Supplementary Fig.
2 ) and should be considered once a stable set of reference genomes has been selected by the community.
As such, a computationally lightweight and easily updatable pangenome-informed genotyping approach is needed.
This problem can be tackled with allele-to-sequence dictionaries for genomic regions of interest 34 , 35 , whereby exact matches of diagnostic short sequences (here, 80 bp k -mers) are counted for each putatively causal allele.
This k -mer-based genotyping approach of a priori known alleles is agnostic to the genomic complexities that can cause misalignment and false discovery of single-reference-based variants, thereby accelerating trait discovery 33 .
To illustrate the power of k -mer-based genotyping to detect complex structural variation, we explored SHATTERING1 ( SH1 ), a YABBY transcription factor that causes loss-of-seed-shattering and is associated with broad geographical divergence and possibly multiple independent domestication events in sorghum 36 , 37 .
Initial exploration of this locus in our pangenome graph revealed known variants that distinguish three haplotypes.
The first, sh1 -1 ( sh1 BTx623-like , Sobic.001G152901), is typical of BTx623 and represents the shortest sequence owing to a known 2,259 bp deletion relative to the undomesticated sequence.
The second, sh1 -2 ( sh1 SC 265-like , SbPI156178.01G139400), is the most common haplotype and has a putative splice modifier.
And the third, sh1 -3 ( sh1 RTx430-like , SbiRTX430.01G157400), exhibits many putatively functional 36 nonsynonymous SNPs.
Our exploration of these haplotypes also revealed a previously undocumented 7,856 bp insertion found only in sh1 -3 and includes an identical 2,161 bp segmental duplicate (Fig.
3a ).
Despite substantial previous efforts to clone and explore SH1 (refs.
36 , 37 ), to our knowledge, this major insertion and untypable (through single-reference short-read alignment) exact duplicate were unknown before our de novo assembly of RTx430 and other pangenome members.
Fig.
3: Deep divergence between domestication locus haplotypes.
a , Simplified ‘sequence tube map’ showing only large indel variants (≥1 kb) between genomes and dot plots across the three sh1 haplotypes.
Representatives of each of the haplotypes were aligned to the longest sh1 -3 haplotype and 80-mer alignments are shown in the dot plots, revealing an insertion relative to sh1 -1 and an exact segmental duplicate in sh1 -3.
The RTx430 coordinates of the SVs in the tube map are reported below the vertical dotted lines.
Positions of uninformative (dark grey), non-unique (light grey) and diagnostic (larger points coloured by haplotype assignments) k -mers are shown.
b , The 1,984 diversity panel re-sequencing libraries coloured by most representative (highest proportion) sh1 haplotype assignment in genome-wide principal component (PC1 and PC2) space (Extended Data Fig.
1 ).
sh1 haplotype proportions for georeferenced African members of the diversity panel (top: pies on the map) and libraries of unique genotypes (bottom: bar chart) illustrate previously observed strong spatial and genome-wide population genetic structure.
Full size image
a , Simplified ‘sequence tube map’ showing only large indel variants (≥1 kb) between genomes and dot plots across the three sh1 haplotypes.
Representatives of each of the haplotypes were aligned to the longest sh1 -3 haplotype and 80-mer alignments are shown in the dot plots, revealing an insertion relative to sh1 -1 and an exact segmental duplicate in sh1 -3.
The RTx430 coordinates of the SVs in the tube map are reported below the vertical dotted lines.
Positions of uninformative (dark grey), non-unique (light grey) and diagnostic (larger points coloured by haplotype assignments) k -mers are shown.
b , The 1,984 diversity panel re-sequencing libraries coloured by most representative (highest proportion) sh1 haplotype assignment in genome-wide principal component (PC1 and PC2) space (Extended Data Fig.
1 ).
sh1 haplotype proportions for georeferenced African members of the diversity panel (top: pies on the map) and libraries of unique genotypes (bottom: bar chart) illustrate previously observed strong spatial and genome-wide population genetic structure.
Diagnostic k -mers that distinguished these three haplotypes were distributed across the entire interval, both at the indel breakpoints and across linked variants (Fig.
3a ), which enabled us to classify all short-read re-sequencing panel members by identity to the three sh1 haplotypes.
Previous work 37 demonstrated that the three non-shattering alleles are most similar to functional SH1 alleles in wild sorghum populations in Tanzania ( sh1 -1), Nigeria ( sh1 -2) and Kenya ( sh1 -3), which led to the hypothesis that independent domestication events in these local regions produced the extant geographical structure and possible divergence among botanical types.
Our assignment of the most representative sh1 allele for a set of 414 georeferenced traditional local varieties supports this claim: sh1 alleles were highly structured by both genetic subpopulation ( χ 2 16 , n 409 = 406.14, P < 0.001; Fig.
3b ) and botanical type ( χ 2 8 , n 474 = 230.32, P < 0.001; Extended Data Fig.
3d ).
For example, sh1 -1 was most abundant among Kafir botanical types (61.1%), whereas sh1 -2 was most abundant among Bicolor (60.0%), Durra (89.0%) and Guinea (82.8%) types.
Finally, sh1 -3 was most abundant among Caudatum (68.2%) botanical types.
These associations between sh1 alleles and botanical types seemed to be strongly associated with continent-wide climate variation.
For example, high temperature seasonality, low April–June photosynthetically active radiation and wet warm seasons clearly distinguished sh1 -1 from the two more common haplotypes (Extended Data Fig.
3d ).
However, the associations between sh1 alleles, genetic subpopulations and botanical types were not fixed.
This finding indicates that introgression or ancient incomplete lineage sorting of sh1 alleles occurred between even the most isolated gene pools 22 , 38 and is potentially related to local consumer and grower preference among botanical types.
Therefore, additional analyses of genetic diversity as a function of gene flow and climate are needed to learn more about the history of sorghum and to detect signatures of adaptation across and in distinct sorghum gene pools.
Climate adaptation and migration balance
Diversity across our pangenome reference revealed ancient divergence between sh1 haplotypes and evidence of multiple independent domestication events 36 , 37 , even though the global sorghum gene pool is only moderately differentiated ( k 10 subpopulation F ST = 0.192; Supplementary Note 4 and Supplementary Fig.
3 ).
This moderate population stratification probably reflects both natural selection and farmer and consumer preferences 1 , 5 and is shaped by the complex interplay among ancient domestication alleles, local adaptation, reproductive barriers, high-diversity cultivation methods and gene flow 5 , 39 .
Thus, contemporary decentralized breeding networks require a broad-based understanding of the patterns and processes that shape local adaptation and migration across the landscape.
We tested for landscape genetic signatures of adaptation, migration and gene flow by analysing spatial patterns of genetic connectivity (sharing of alleles) among 433 georeferenced African and southern Arabian cultivars through two related approaches.
The first method was multilocus wavelet genetic dissimilarity 40 , which models allele frequency changes and identifies spatial scales at which genetic similarity is higher or lower than expected under panmixia (random mating).
The second approach was estimated effective migration surfaces 41 , 42 , a graph-based method that describes spatial patterns and local rates of gene flow across the landscape at fixed spatial scales.
Both models clearly demonstrated the impact of human-mediated gene flow on sorghum diversity.
For example, sorghum allelic dissimilarity deviated from panmixia at 125 km, 17 times more distant than highly structured selfing populations of wild Arabidopsis thaliana (7 km) 40 , but similar to that of the highly outcrossing African crop pearl millet (140 km) (Fig.
4a and Extended Data Fig.
3 ).
Despite the pervasive outcrossing of pearl millet, allele sharing among sorghum populations exceeded pearl millet at larger and continental geographical scales, thereby demonstrating the importance of long-distance and probable human-mediated gene flow.
Furthermore, effective migration surfaces of sorghum revealed local areas of low gene flow in the Ethiopian highlands and in the western and central Sahel, and continent-wide gradients along the two African continental axes that have been proposed to drive sorghum adaptive differentiation (Fig.
4b ).
Consistent with locally low rates of effective migration, both the Ethiopian highlands and Sahel seemed to have experienced secondary contact among domestication haplotypes (Fig.
3b ) and are locations that have been previously shown to have local adaptation across steep environmental gradients in sorghum and other crops 14 , 15 , 36 , 37 .
Fig.
4: Spatial and climatic context of gene flow.
a , Comparison of the geographical scale of allele frequency change (multilocus wavelet allelic dissimilarity) in Arabidopsis (green), sorghum (purple) and pearl millet (black).
Dashed lines are mean observed differentiations, and coloured polygons indicate null (panmixia) range based on permutation tests.
Geographical distance (km) is log-scaled on the x axis.
As dissimilarity is not necessarily comparable among species and may be affected by sample size and marker diversity, the y axis is scaled so that the range of observations are equivalent across species (see Extended Data Fig.
3 for raw allelic dissimilarity values).
b , Estimated effective migration surfaces among sorghum populations in north–south (left) and east–west (right) corridors, which have each been shown to have adaptive variation in cultivated sorghum.
High gene flow (blue) and low gene flow (orange) are indicated by coloured edges in the wire-mesh graph.
Graph grid resolution corresponds to cell spacing (distance between mid-points of adjacent cells) of 110 km.
c , Estimated effective migration surfaces between low-drought and high-drought prevalence sorghum populations in the east–west corridor.
d , Comparison of the geographical scale of allele frequency change between low-drought prevalence and high-drought prevalence sorghum populations; means of each grouping are shown in white-bordered dashed lines.
Source data Full size image
a , Comparison of the geographical scale of allele frequency change (multilocus wavelet allelic dissimilarity) in Arabidopsis (green), sorghum (purple) and pearl millet (black).
Dashed lines are mean observed differentiations, and coloured polygons indicate null (panmixia) range based on permutation tests.
Geographical distance (km) is log-scaled on the x axis.
As dissimilarity is not necessarily comparable among species and may be affected by sample size and marker diversity, the y axis is scaled so that the range of observations are equivalent across species (see Extended Data Fig.
3 for raw allelic dissimilarity values).
b , Estimated effective migration surfaces among sorghum populations in north–south (left) and east–west (right) corridors, which have each been shown to have adaptive variation in cultivated sorghum.
High gene flow (blue) and low gene flow (orange) are indicated by coloured edges in the wire-mesh graph.
Graph grid resolution corresponds to cell spacing (distance between mid-points of adjacent cells) of 110 km.
c , Estimated effective migration surfaces between low-drought and high-drought prevalence sorghum populations in the east–west corridor.
d , Comparison of the geographical scale of allele frequency change between low-drought prevalence and high-drought prevalence sorghum populations; means of each grouping are shown in white-bordered dashed lines.
These analyses also implicate drought as a major force in historical and contemporary adaptations of sorghum 2 , 4 , 43 .
To test the hypothesis that drought adaptation affects the geographical scale of gene flow, we binned accessions by those originating from high-drought ( n = 258) and low-drought ( n = 175) prevalence populations (Fig.
4c and Extended Data Fig.
3 ).
Drought status was broadly different among admixture ( k = 10) subpopulations ( χ 2 9 , n 431 = 82.664, P < 0.001), and we observed different patterns of allele sharing in drought classes.
For example, accessions from high-drought prevalence populations had lower allelic dissimilarity than those from less drought-prone populations (Fig.
4d ).
This result suggests that alleles were more likely to move between spatially disjunct populations experiencing drought.
This pattern was robust across the continent, with similar increases in drought-adapted allele sharing in each of the six nonoverlapping geographical regions analysed (Extended Data Fig.
3 ).
However, migration patterns in the Sahel complicated this finding.
At this location, our observation of generally lower effective migration rates persisted only among cultivars from northern, more drought-prone habitats.
Conversely, effective migration among cultivars from less drought-prone southern Sahel habitats was higher than average (Fig.
4c and Extended Data Fig.
3 ).
Dhurrin biosynthesis in pangenomic variants
Although continent-scale gradients of abiotic stress intensity have clearly shaped sorghum diversity, local agricultural productivity is often equally affected by adaptation to complex interactions among pests, pathogens and nutrient availability.
Drought-adaptive and pest-adaptive responses (for example, stomatal regulation and structural leaf defence) are obvious individual targets for breeders and natural selection alike, whereas some secondary metabolites offer pleiotropic resistance to both stresses.
For example, the cyanogenic glucoside dhurrin not only provides resistance to chewing insect herbivory via cyanide release 48 but is also considered to improve constitutive dehydration avoidance 49 .
As such, dhurrin is an effective bridge between physiological, biochemical and broader phenotypes such as plant vigour and WUE that influence field performance.
We first sought to define individual loci associated with the biosynthesis and catabolism of dhurrin by phenotyping members of the diversity panel for seedling dhurrin content ( n = 175) and hydrogen cyanide potential (HCNp, n = 367).
Contrary to expectations of a single function of dhurrin for pest defence through HCNp, the two traits were uncorrelated at the seedling stage (3–4-leaf stage; Pearson’s r = 0.075, P = 0.33) and only weakly genetically correlated later in vegetative growth (7–8-leaf stage; Pearson’s r = 0.16, P < 0.05).
Accordingly, none of the strongest genome-wide association (GWAS) peaks for dhurrin (Fig.
5a ) and HCNp (Extended Data Fig.
5a and Supplementary Data 9 – 11 ) were physically proximate, which potentiates a functional role of modifications in dhurrin levels outside pest defence.
Dhurrin biosynthesis and transport are governed in part by a five-gene biosynthetic gene cluster (BGC) on chromosome 1 (ref.
50 ).
Correspondingly, one of the strongest and the densest associations in our dhurrin GWAS resided in the approximately 175 kb BGC (Fig.
5b ).
Although the majority of significant variants were in intergenic regions, an alanine-to-valine missense mutation at 1,167,702 bp in CYP79A1 (Sobic.001G012300) was predicted to increase the binding affinity of cytochrome P450 for tyrosine, the substrate for the first committed step of dhurrin biosynthesis 51 (Extended Data Fig.
5b,c ).
There were also several strong regulatory candidate variants, including a 2 bp CT deletion at 1,185,567 bp that disrupted predicted binding sites in an accessible chromatin region for abscisic-acid-responsive transcription factors (ABF3 and ABF4; Supplementary Table 3 ).
As functional sequence variation in BGCs can affect traits both directly and epistatically through modification of the pathway (for example, abundance of precursors) 21 , it is important to consider the BGC as a whole when designing genome-enabled approaches to improve drought and pest adaptation through dhurrin production.
Consistent with the co-inheritance of, and probable correlated selection on, the BGC, we observed strong linkage disequilibrium (LD) blocks in which all but one significant variant fell into three linked clusters (Fig.
5b ).
These four groups of markers represented most of the unlinked variation in the region that is associated with dhurrin concentration.
Combined, the total number of non-reference BTx623 alleles across these four blocks (Fig.
5c ) and two additional sites with higher missing data (Extended Data Fig.
5d ) were highly predictive of dhurrin concentration owing to a marginally significant additive (linear model coefficient estimate t = –2.18, P = 0.03) and highly significant quadratic ( t = 4.97, P < 0.0001) effect on untransformed dhurrin levels.
This observation indicates that linked epistatic interactions or other nonlinear effects between adjacent loci result in genotypic combinations with highly increased dhurrin levels.
Breeding efforts at linked loci such as the BGC must also consider the combined effects of the interval as a whole.
Such an objective is ideally suited to our pangenome k -mer genotyping approach.
Pangenome graph haplotype clustering revealed that 32 out of the 33 reference pangenome members fell neatly into four tight k -mer identity clusters that distinguished samples by previously typed short variants (Fig.
5c ) and several previously unknown large intergenic indels (Fig.
5d ).
The unclustered member IRAT204 was intermediate and exhibited signatures of recombination between the ‘high’ and ‘mid’ clusters.
This four-level clustering explained 3% more variance in seedling dhurrin content, a 15% better fit than the most predictive individual variant.
Notably, the pangenome-based clustering revealed previously unobserved trait and climate associations.
Diversity panel members with the highest proportion of k -mers diagnostic to the ‘high’ and ‘mid’ dhurrin BGC clusters produced the most dhurrin and were georeferenced to localities with lower annual precipitation (Kruskal–Wallis rank sum H = 41.01, P < 0.0001; Fig.
5e,f ).
This effect was even stronger at local scales, especially among samples from west Africa ( H = 77.48, P < 0.0001; Fig.
5f ).
Thus, the BGC seems to be a metabolic hub that underlies variations in pest resistance and is probably structured by adaptation to moisture availability across continental and local geographical scales.
Expanding food security and economic prosperity under rapidly changing environmental stresses will require transformative advances in global crop improvement speed and efficacy 52 .
Here we introduced a pangenomic resource and described our efforts to integrate pangenome-enabled variant discovery to define targets of historical environmental adaptation and to accelerate breeding decisions in sorghum.
By pairing each reference genome with standardized trait data, we created a functional framework that links genomic diversity to observable agronomic performance, thereby providing species-wide biological context rather than population-level inference.
The k -mer genotyping method described in this work has been applied to other complex loci with structural variation (for example, LGS1 (ref.
33 ) and Resistance to Melanaphis sorghi 1 ( RMES1 ) 53 ) and used for marker development, which illustrates both the utility of this approach and, more broadly, the value of translating patterns of diversity in pangenome references to diverse germplasm for crop improvement.
In addition to better characterizing large SVs, our pangenome reference facilitates information transfer between genotypes used in breeding and those that are more amenable to laboratory experimentation.
This advantage is especially important in sorghum, which is highly recalcitrant to genome-editing methods.
Indeed, so far, none of the major breeding lines can be efficiently edited and only two genotypes (RTx430 and P898012) are considered readily transformable without morphogenic regulator technology.
Our pangenome resource bridges this gap by effectively reconstructing the putative functional alleles in breeding pedigrees and the orthologous sequence to target in transformable varieties.
Consequently, it will pave the way for accelerated pangenome-enabled traditional breeding and genome editing of locally adapted alleles across global sorghum germplasm.
Plant material preparation and nucleic acid extractions
Young seedlings were grown in flat pans under healthy, pest-free and disease-free conditions until the first fully developed leaves emerged.
To optimize DNA yield and quality, seedlings were dark-treated for 24–30 h under moist conditions.
Tissue was collected by hand in small batches, cutting half an inch above the soil surface and immediately flash-freezing in liquid nitrogen within 1 min of excision.
Approximately 50 g of tissue was collected in this manner, stored in pre-labelled freezer-quality ziplock bags at –80 °C and kept frozen until extraction.
DNA was extracted from young tissue using a previously published protocol 54 with minor modifications.
Flash-frozen young leaves were ground to a fine powder in a frozen mortar with liquid nitrogen followed by gentle extraction in 2% CTAB buffer (including proteinase K, PVP-40 and β-mercaptoethanol) for 30 min to 1 h at 50 °C.
After centrifugation, the supernatant was gently extracted twice with 24:1 chloroform and isoamyl alcohol.
The aqueous phase was transferred to a new tube and one-tenth volume 3 M sodium acetate was added, gently mixed and DNA was precipitated with isopropanol.
DNA precipitate was collected by centrifugation, washed with 70% ethanol, air dried for 5–10 min and dissolved thoroughly in an elution buffer at room temperature followed by RNAse treatment.
DNA purity was measured with Nanodrop, and the DNA concentration was measured using a Qubit HS kit (Invitrogen).
DNA size was validated using a Femto Pulse system (Agilent).
We also grew three biological replicates of the reference genotypes (see Supplementary Data 12 for phenotype data and Supplementary Data 13 for original photographs without the backgrounds removed) and sampled them at key developmental stages and organ types for transcriptome assembly and genome annotation (sample and data collection were performed by the team at Donald Danforth Plant Science Center in 2020).
For the 29 diverse pangenome reference members and BTx623 (excluding, BTx642, Wray and RTx430), tissues were collected from greenhouse-grown sorghum plants maintained under controlled environmental conditions.
For each replicate, tissues from multiple plants were pooled when individual tissue quantity was insufficient.
Six tissue types were represented in the samples.
(1) Approximately half-inch sections of stem tissue were excised from each internode and nodal region of the same stem at 60 days after planting (DAP), immediately frozen in liquid nitrogen and pooled to generate composite stem samples.
(2) At 28 DAP, the youngest fully expanded leaf was collected and subdivided into tip, base, midrib and sheath segments, which were pooled to represent the composite leaf sample.
(3) Entire primary and crown root systems were harvested at 28 DAP, gently rinsed with water to remove adhering soil and pooled per replicate.
(4) Developing inflorescences were collected at both heading and anthesis to capture pre-pollination and post-pollination reproductive stages.
Whole inflorescences were excised, flash-frozen and pooled across plants.
(5) Developing grain was sampled at 5 DAP, 20 DAP and at the black layer stage to represent early, mid and late seed development, respectively.
(6) Whole seedlings were collected at 12 DAP under well-watered (100% field capacity (FC)) and water-stressed (45% FC) conditions.
Water stress was initiated 5 DAP, and both shoot and root tissues were harvested and pooled at 12 DAP.
In addition to these developmental-stage collections, time-course sampling was conducted for two representative genotypes, PI660565 and PI276816, to capture early vegetative dynamics.
For each line, three biological replicates were collected at 10, 17, 21, 25 and 31 DAP.
At each time point, leaf and stem tissues were collected following the same procedures described above.
Immediately after collection, all tissues were flash-frozen in liquid nitrogen and stored at –80 °C until RNA extraction.
Tissue collection for Wray and for BTx642 and RTx430 (as previously detailed 26 ) generally followed the above protocol, but with some differences in the types of tissues.
Long-read sequencing technology, as used in the construction of our pangenome reference assemblies, has improved the resolution of repetitive regions and closed gaps in other complex genomes 55 , 56 .
Although the quality and contiguity of our pangenome assemblies are consistent, the input data depth, sequencing technology, bioinformatic methods available and sequencing read characteristics necessitated some differences in computational genome assembly strategy.
Here we provide a summary of the basic methods (see Supplementary Note 2 for a full description of genome assembly methods for each reference genome).
The majority of the pangenome sequences were generated by assembling PACBIO CLR reads with CANU (v.1.8) 57 , whereas five accessions (PI156178, RTx430, BTx642 and PI565121) were assembled with MECAT (v.1.4) 58 , and one accession (Wray) was generated using PACBIO HiFi data assembled with HiFiAsm+HIC (v.16.1) 59 .
All sequencing was conducted at the HudsonAlpha Institute for Biotechnology and the Department of Energy (DOE) Joint Genome Institute.
All assemblies were subsequently polished using ARROW (v.2.1) 60 , except BTx623 and Wray, which were polished using RACON (v.1.14) 61 .
A combination of 32,400 syntenic markers (unique, non-repetitive, nonoverlapping 1.0-kb sequences from BTx642) and 12,641 annotated genes from BTx623 were used to identify misjoins in the assembly.
Contigs were then ordered, oriented and assembled into ten chromosomes after examining syntenic marker and gene alignment positions on the polished assemblies.
Each chromosome join was padded with 10,000 Ns.
Contigs terminating in significant telomeric sequence were identified using the (TTTAGGG) n repeat, and care was taken to make sure that they were properly oriented in the production assembly.
Chromosomes were numbered using BTx623 V3 naming convention.
The remaining scaffolds were screened against bacterial proteins, organelle sequences, and GenBank non-redundant datasets and removed if found to be a contaminant.
Chloroplast and mitochondrial genomes were generated using the OatK pipeline (v.1.0) 62 .
Homozygous SNPs and indels were corrected to match the consensus call from Illumina fragment reads (2 × 150, 400 bp insert) by aligning the reads using bwa-mem (v.0.7.17-r1188) 63 and identifying homozygous SNPs and indels with the UnifiedGenotyper tool in GATK (v.3.6-0-g89b7209) 64 .
Genome annotation was performed using the pipeline developed by the DOE Joint Genome Institute and Phytozome 65 .
As methodological considerations can substantially affect estimates of protein-coding gene PAVs 30 , each pangenome member underwent two rounds of gene prediction.
In the first round, gene models were independently predicted for each genome followed by the propagation of these predictions across the entire pangenome in the second round.
In the initial round, transcript assemblies were generated from 2 × 150 bp stranded paired-end Illumina RNA sequencing reads (Supplementary Table 2 ) using PERTRAN (as previously described in detail 66 ) for genome-guided assembly via GSNAP (v.2013-09-30) 67 , followed by splice graph construction, alignment validation and correction.
PacBio Iso-Seq CCS reads, when available, were corrected and collapsed using a GMAP-based pipeline 67 to refine alignments, correct splice junctions and cluster alignments when their intron (or introns) matches if spliced or 95% similar if it was a single exon.
Final transcript assemblies were produced using PASA (v.2.0.2) 68 , integrating RNA sequencing assemblies, corrected CCS reads and Sanger ESTs.
A species-specific repeat library was built from BTx623 V3 using RepeatModeler (v.open1.0.11) 69 .
Repeats were functionally analysed with InterProScan (v.5.47-82.0) 70 , incorporating Pfam 71 and PANTHER 72 databases, and those with significant hits to protein-coding domains were excluded.
Genomes were soft-masked using RepeatMasker (v.open1.0.11) 69 with the curated repeat library.
Gene loci were identified using transcript assembly and EXONERATE (v.2.4.0) 73 alignments of proteins from Arabidopsis , soybean, poplar, Aquilegia , grape, rice, Setaria viridis , Brachypodium , Panicum hallii , pineapple, Acorus americanus and Swiss-Prot (2020_06) to the repeat soft-masked genomes, with up to 2,000-bp extension on both ends unless extending into another locus on the same strand.
Gene models were predicted using FGENESH+ (v.3.1.0) 74 , FGENESH_EST, EXONERATE, PASA assembly ORFs and AUGUSTUS (v.3.1.0) 75 trained on high-confidence PASA assembly open reading frames with intron hints from RNA sequencing alignments.
Candidate models were selected based on EST and protein support and penalized for repeat overlaps.
PASA refinement added UTRs, corrected splicing and incorporated alternative isoforms.
Cscore (BLASTP score ratio and protein coverage) was computed for PASA-refined proteins.
Transcripts were retained if Cscore and coverage was ≥0.5 or supported by ESTs; those with >20% CDS-repeat overlap required a Cscore ≥ 0.9 and ≥ 70% coverage.
Models with >30% TE domain overlap (Pfam) were excluded.
In the second round, each genome was hard-masked with its high-confidence gene models (supported by transcriptome and homology evidence).
BLASTX and EXONERATE were used to predict new gene models by projecting high-confidence models from other pangenome members onto the hard-masked assemblies.
Predicted models were retained if they showed stronger homology support than existing models and were not contradicted by transcript evidence, or if no first-round model existed at that locus.
Incomplete gene models, which had low homology support without full transcriptome support, or short single exon genes (<300 bp CDS) without protein domain or good expression were manually excluded.
Pangenome graph construction and exploration
A pangenome graph was constructed for each chromosome using Minigraph-Cactus and default settings (v.2.9.3) 76 with the BTx623 V5 genome as the primary reference.
Clipped chromosome graphs were merged and prepared for visualization using vg (v.1.59.0) 77 .
We used ODGI (v.0.9.0) 78 to inspect representation of each genome across the graph and sequenceTubeMap (v.0.1.0) 79 to visualize variation in genomic regions of interest.
For the dhurrin BGC and SH1 loci, we further processed this graph with vg and vcfwave (vcflib v.1.0.10) 80 to reduce allelic complexity and retain only variants ≥5 kb and ≥1 kb in length, respectively.
Gene families were calculated with OrthoFinder (v.2.5.5) 81 and parsed with GENESPACE (v.1.3.1) 82 to create saturation curves.
Synteny maps were created using DEEPSPACE (v.0.1.0, github.com/jtlovell/DEEPSPACE ), which aligns and tracks the positions of windowed sequence alignments and quickly visualizes large-scale SVs.
SyRI (v1.6.3) 83 was used for pairwise variant detection from whole-genome alignments via minimap2 (v.2.22-r1101) 84 .
Pangenome graph saturation curves were calculated with Panacus 85 .
DNA re-sequencing and variant detection
A total of 2,145 diversity samples ( n = 1,984 unique genotypes + redundant + polishing) were re-sequenced at a median coverage of 43.39× (range 1.96–364.18) (Supplementary Data 3 ).
The samples were sequenced using Illumina HiSeq X10 and Illumina NovaSeq 6000 paired-end sequencing (2 × 150 bp) at the HudsonAlpha Institute for Biotechnology and the Joint Genome Institute.
To account for different library sizes, reads were downsampled to ≤50× coverage.
SNPs and short indels were called by aligning Illumina reads to the BTx623 V5 reference with bwa mem 63 .
The resulting .bam file was filtered for duplicates using Picard ( http://broadinstitute.github.io/picard ) and realigned around indels using GATK (v.3.056) 64 .
Multisample SNP calling was done using SAMtools (v.1.7) mpileup 86 and Varscan (v.2.4.089) 87 with a minimum coverage of eight and a minimum alternate allele count of four.
Genotypes were called using a binomial test.
Variants within 25 bp of a 24-mer repeat were removed from further analyses.
For most analyses, only SNPs with minor allele frequencies (MAFs) of >0.001 were retained, which resulted in 36,061,325 SNPs at a coverage depth between 8× and 500×.
Phasing was performed using SHAPEIT (v.3) 88 .
To project patterns of observable diversity in the pangenome assemblies to the broader whole-genome resequencing panel, we applied k -mer-based genotyping to SH1 and the dhurrin BGC locus.
In brief, the k -mer genotyping methodology extracts ancestry-informative k -mers 34 , 35 for genomic regions of interest and counts exact matches of these k -mers in Illumina sequencing libraries.
In this instance we used 80-mers that were globally single-copy (but could be duplicated locally) within individual references and absent in at least one member of the pangenome.
This methodology is analogous to a k -mer-based GWAS 89 , but instead of testing for genome-wide associations using accession-derived k -mer PAV, it aggregates k -mers from pangenome references that are unique to and diagnostic of a haplotype in a predefined region of interest (ROI).
The pipeline proceeded as follows.
First, each reference genome was individually converted into k -mers; each k -mer and its frequency in the assembly were stored in a hash table.
Any non-adjacent multicopy k -mers were flagged to be ignored during downstream analyses.
Second, individual reference hashes were combined in a single hash (‘main hash’), and the flags were updated so that only single copy k -mers among all considered references were used downstream (for example, a single-copy k -mer in reference A was ignored if it was multicopy in reference B).
To isolate k -mers that were useful for genotyping a ROI (for example, the SH1 or dhurrin BGC loci), the orthologous region from each reference was extracted (from either from the pangenome graph or best alignment bounds) using minimap2 (ref.
84 ).
Steps 1 and 2 above were repeated on the subsetted fasta files, which generated a combined subsetted hash that contained local single-copy sequences from all ROI orthologues across the pangenome.
Third, to ensure that the ROI local single-copy markers did not occur elsewhere in the genome, which would prevent their use as a genotyping marker, the combined ROI hash was intersected with the main genome hash, and flags were updated so that any non-single copy k -mers were ignored when genotyping (‘genotyping hash’).
The total number of diagnostic k -mers (globally single copy and not common to all pangenome reference members) used in the genotyping approach was 6,533 for the SH1 locus and 139,866 for the dhurrin BGC locus.
Fourth, Illumina libraries were genotyped by searching for all k -mers in the genotyping hash and counting their frequencies.
At this point, counts were either merged into a single matrix for de novo clustering (for example, dhurrin BGC locus), or binned based on a priori clusters (for example, SH1 locus) based on the local haplotype structure at the region of interest.
De novo clustering as used for k -mer genotyping the dhurrin BGC locus operated on a binary matrix of n × m ( k -mer presence/absence × Illumina libraries), where k -mer frequencies greater than one were considered present.
Clusters were visualized in principal component space and coloured by clustering as defined by partitioning around medoids (PAMs) of pairwise Jaccard distances among reference libraries.
A priori clustering was used for k -mer genotyping the SH1 locus as the per cent match to diagnostic k -mers in each bin.
Illumina library haplotypes were then assigned to a SH1 cluster based on majority match to haplotype bins.
To generate the dot plots in Fig.
3 , the position of SH1 was queried from the pangenome graph by ‘vg find‘, returning intervals the following nodes in the native genome coordinate systems on chromosome 1: BTx623: 12,398,577–12,400,453 bp, 12,400,454–12,401,958 bp, 12,401,959–12,404,976 bp; MN2014: 12,177,214–12,179,090 bp, 12,179,091–12,181,356 bp, 12,181,357–12,182,863 bp, 12,182,864–12,185,897 bp; RTx430: 12,088,419–12,090,291 bp, 12,090,292–12,092,556 bp, 12,092,557–12,094,061 bp, 12,094,062–12,101,908 bp, 12,101,909–12,104,942 bp.
These contiguous sequence blocks (BTx623 12,398,577–12,404,976 bp; MN2014: 12,177,214–12,185,897 bp; RTx430: 12,088,419–12,104,942 bp) were converted to 80-mers and aligned to the RTx430 genomic interval with minimap2 (parameters = -r80,800 -O6,26 -e2,1 -B4 -f0.0001 -p0.5 -N100 -k19 -w12 -g160 -F800 -t8 -A1 -U50,500 -no-long-join -rmq=no -n1 -m0 -frag=yes), retaining the primary (mapq = 60) hits or those hits within 1 bp of the number of mismatches in the primary alignment.
The aligned k -mers were binned into those among the diagnostic 80-mers (see above) or those that are not diagnostic and coloured according to the genome to which they are unique.
Diversity panel source material
The sorghum diversity panel used in this study comprised 1,984 unique accessions sourced from structured diversity panels and curated regional collections.
These lines spanned Africa, Asia and the Americas and represented both landraces and breeding lines, thereby providing broad coverage of genetic, phenotypic and geographic variation of sorghum and 10 discrete genetic resources or populations.
(1) CASP: 82 lines from a 648-line association panel previously described 90 , which was developed for trait mapping under field and drought conditions.
(2) Biomass Association Panel (BAP): 390 accessions representing biomass-related diversity across sorghum botanical types and global regions 91 .
The panel is widely used in bioenergy trait mapping, and 375 of the BAP accessions are included in this study.
(3) Sorghum Association Panel (SAP): 377 genotyped accessions developed by USDA-ARS and collaborators 92 , which represents all five major sorghum races and is widely used in GWAS.
A total of 328 SAP accessions are included in this study.
(4) TERRA MEPP: represents a wide range of traits relevant to field-based high-throughput phenotyping.
These lines were selected to enable trait discovery under environmental variation 93 .
(5) Purdue Inbred Calibration Set: a set of accessions with which emphasize improving traits related to drought, cold tolerance, striga resistance, nutritional quality and biomass potential.
These lines have played a central part in UAV-based remote-sensing studies that integrate hyperspectral and LiDAR data with marker profiles, which underscores their relevance in phenomics–genomics research 94 , 95 .
(6) Georeferenced Panel: 529 lines including 166 accessions from the BAP, 133 African accessions from the WRS and an additional 230 accessions from NPGS-GRIN, selected to maximize spatial sampling.
Georeference coordinates were sourced from Genesys using ICRISAT IDs (IS numbers) and cross-referenced with NPGS-GRIN IDs (PI numbers) for GRIN accessions.
(7) World Reference Set (WRS): 383 accessions 96 , which capture broad allelic diversity, including all five major cultivated races and their intermediates.
The set reflects extensive geographical coverage, spanning central and eastern Africa to Western Africa, South Asia and East Asia, with significant representation in domesticated morphotypes rather than strict race categories.
Our study includes 381 of these 383 accessions, thereby preserving nearly the full scope of genetic and geographic diversity defined therein.
(8) West African Sorghum Association Panel (WASAP): a 756-member diversity panel contributed by national programmes in Mali, Senegal, Togo and Niger to represent West African landraces and regionally adapted breeding lines.
(9) Ethiopian Institute of Agricultural Research (EIAR): a regional collection developed and curated by EIAR to capture the exceptional diversity of Ethiopian highland sorghum.
These accessions include seed-increased landraces and genotyped lines selected for local adaptation.
(10) BC-NAM Parents: 37 founder accessions from a West and Central African Backcross Nested Association Mapping (WCA-BCNAM) population, as previously partially detailed 97 .
These lines were selected for adaptation traits and crossed with three elite recurrent parents from the region, yielding 3,901 BC-NAM lines.
The founders were chosen to maximize genetic diversity in plant height, flowering time and environmental response across agro-ecologies varying in photoperiod, rainfall, temperature and fertility 97 .
This panel structure reflects global efforts to unite diverse germplasm for climate-aware trait discovery and genomic breeding.
Passport data collection and curation
Passport data—including botanical classification, breeding improvement status, country of origin and geographical coordinates—were compiled from multiple sources: the GRIN-Global database, Genesis Global, the ICRISAT Sorghum Passport dataset and previously published studies 93 , 98 .
Records were cross-referenced to identify and resolve discrepancies across datasets.
In cases where inconsistencies could not be reconciled, both conflicting metadata entries were retained and included in the final metadata.
We applied ADMIXTURE (v.1.3.0) 99 using default settings to n = 1,984 unique genotypes of non-duplicated sequencing libraries.
To generate an approximately independent set of markers, we removed variants with MAF < 0.05, more than 50% missingness, and applied LD-based pruning with a window size of 50 SNPs, step size of 5 SNPs and a r 2 threshold of 0.5 in PLINK (v.1.90b6.12) 100 .
After filtering, we randomly selected 100,000 SNPs to estimate the optimal K of ancestral groups (optimal K ≈ 10, see Supplementary Note 4 and Supplementary Fig.
3 for details).
For the remainder of population genetic analyses, we retained n = 433 samples georeferenced to the African continent that were designated as landraces, cultivars or wild strains and missed a genotype call in <15% sites.
To select sites that captured dynamics of population structure, we selected putatively neutral and unlinked variants using bcftools (v.1.9) 101 with the following filtering parameters: invariant sites and sites with strand bias in variant-supporting reads (>90%) were removed.
We required all retained variants to be synonymous, have a genotype call in at least 95% of individuals sampled and a MAF ≥ 0.01.
Filtered genotypes were imputed with beagle (v5.4) 102 and variant effects annotated with SnpEff (v5.1d) 103 .
We further LD-pruned the remaining SNPs in PLINK using 50-SNP windows with a variance inflation factor (VIF) of 2 (VIF = 1/(1 – R 2 ), where R 2 is the multiple correlation coefficient for a SNP being regressed on all other SNPs simultaneously, PLINK documentation), resulting in a set of 33,823 neutral LD-pruned SNPs.
We used these 33,823 SNPs to estimate effective migration surfaces in fast estimation of effective migration surfaces (FEEMS 41 ).
We ran separate analyses for the African continental axes (north–south and east–west).
For each independent run of FEEMS we optimized the smoothing regularization parameter ( λ ) and utilized the optimal lambda for each analysis.
Node and edge positions and weights were exported and formatted for plotting in R.
To describe patterns in allele frequency change at neutral sites capturing dynamics of population structure, we again used the set of 33,823 imputed, neutral and LD-pruned SNPs.
Georeferenced samples ( n = 433) were grouped into populations by averaging allele frequencies of samples georeferenced to the same latitude and longitude (rounded to 3 decimal places, around 100 m), which resulted in 343 populations comprising 1 and 13 individuals (mean = 1.26).
We estimated scale-specific genome-wide turnover by modelling the allele frequency change with wavelet transformations at 15 scales, spaced along a log sequence from the 0.1st to 99th percentile of distances between sites: 24 km, 35 km, 49 km, 69 km, 98 km, 139 km, 197 km, 278 km, 394 km, 558 km, 790 km, 1,119 km, 1,584 km, 2,242 km and 3,174 km.
We compared patterns of sorghum wavelet-transformed allele frequencies to previous estimates in A.
thaliana 40 , and to an analysis of published pearl millet genotype data 104 .
Population-averaged pearl millet genotypes 104 were filtered to remove any markers with missing data, which retained 138,948 markers from 174 populations.
We conducted the scale-specific wavelet analysis at 15 scales, spaced along a log sequence from the 0.1st to 99th percentile of distances between sites: 33 km, 44 km, 59 km, 79 km, 105 km, 140 km, 187 km, 249 km, 332 km, 443 km, 591 km, 788 km, 1,051 km, 1,400 km and 1,867km.
Following previously described methods 105 , we also used wavelet transformations of our sorghum genotypes to determine outlier loci with more rapid spatial allele frequency change compared with the overall distribution, which represent loci that may be important in adaptation.
We used 5,615,029 SNPs filtered for MAF > 0.01 and heterozygosity <0.9 (which are probably representative of genotyping errors).
The top 1,000 outliers from each chromosome at each spatial scale were used in downstream analyses, including a gene ontology enrichment using topGO (v.2.42.0), an R Bioconductor package.
Using the same set of SNPs, we identified putative selective sweeps using normalized xpnsl (v.1.3.0) 44 , grouping georeferenced populations into geographical regions using PAM and comparing individuals from high-drought prevalence sites to individuals from low-drought prevalence sites in each of five geographical regions (a sixth region included only high-drought prevalence sites and was excluded).
We retained all sites with normalized scores above the critical threshold and converted runs of sites into blocks.
We then quantified pairwise base-pair overlap of xpnsl extended haplotypes reciprocally between all geographical regions.
We determined whether overlap was greater than expected by chance by comparing the observed overlap to the overlap of permuted genomic blocks, significance testing relative to 1,000 permutations.
We compared the amount of haplotype overlap using paired t -tests for each geographical region.
Assignment of landraces to climate-based clusters
Following previously described methods 106 , the Cycles agroecosystem model was used to characterize variables representative of stage-specific water stress (vegetative and reproductive) for the point of origin for n = 326 sorghum landrace accessions.
Cycles is a process-based multiyear and multispecies agroecosystem model that simulates biophysical and management practices in cropping systems 107 , 108 to simulate crop growth.
All simulations were carried out using Cycles (v.0.13.0; https://github.com/PSUmodeling/Cycles ).
The crop description file defines the physiological and management parameters that control the growth and harvest of crops used in the simulation.
We used the base Cycles sorghumMS parameters from the default crop description file.
The management (operation) file defines the daily management operations to be used in a simulated crop rotation.
We activated conditional planting where Cycles ‘plants’ a simulated crop once certain soil moisture and temperature levels are satisfied in a window of planting dates.
We turned on the automatic nitrogen fertilization option and set planting density to 67% so that stress observed in model outputs was due entirely to climatic factors.
To approximate the climate conditions the sorghum accessions were adapted to, we simulated plant growth using weather data from 1970 to 1989, 10 years before and after the average collection year.
Daily weather files at one-quarter degree resolution that overlapped with at least one of our georeferenced accessions and for years included in the simulation were fetched from the meteorological data source Global Land Data Assimilation System (GLDAS 109 ) using the script LDAS-forcing.py sourced from GitHub ( https://github.com/shiyuning/LDAS-forcing ).
Soil physical parameters describing the average soil characteristics and land-use type of each landrace accession point of origin were obtained from the ISRIC SoilGrids global database 110 via the HydroTerre data system 111 , 112 , 113 .
Using model outputs and for each landrace accession, we extracted integrative environmental variables representative of stress when in the vegetative and reproductive phase.
Accessions were clustered into two groups (low-drought and high-drought prevalence) using k -means clustering implemented in the R stats package.
As additional georeferenced landraces were added to the diversity set ( n = 107), cluster membership was predicted using a feedforward artificial neural network (ANN) implemented in the R package neuralnet (v1.44.2, https://CRAN.R-project.org/package=neuralnet ).
The ANN used bioclimatic predictors from the WorldClim 2.1 dataset (1970–2000) 114 , interpolated at 10 arcmin resolution (around 340 km 2 per pixel).
Input variables included mean temperature of the driest quarter (MTDQ), mean temperature of the warmest quarter (MTWQ), precipitation of the wettest month (PWM), precipitation seasonality (PS), precipitation of the driest quarter (PDQ) and precipitation of the warmest quarter (PWQ).
To evaluate variation in dhurrin concentration and HCNp among diverse sorghum germplasm, we conducted a single time-point phenotypic analysis using accessions sourced from the USDA Germplasm Resources Information Network (GRIN).
All accessions were grown under controlled conditions at the Colorado State University Plant Growth Facilities (HCNp was analysed during first planting in autumn 2022 and second planting in spring 2023; dhurrin was analysed in spring 2023).
A completely randomized block design was implemented with two replicates per accession.
Four plants per accession were sown in 11.4 l pots containing Pro-Mix HP supplemented with one tablespoon of Osmocote fertilizer.
Accessions were evaluated across two separate plantings, each containing two replicate blocks.
In each block, individual plants were grown and subsequently sampled for trait assessment.
Sampling for HCNp was performed at the seedling stage (3–4-leaf stage) and again at the early vegetative stage (7–8-leaf stage), with tissues from the youngest fully expanded leaf collected from each plant.
Dhurrin was estimated at the seedling stage for only a single replicate in one of the plantings.
All plant tissue samples were processed individually and randomly assigned to plates to control for technical variation.
Blocks were nested in plantings to account for potential variation.
Plates were kept on ice during sample collection and stored at –20 °C.
Cyantesmo test strips (Macherey–Nagel) were cut to fit each plate, applied across alternating rows and sealed with Axygen microplate film (Corning).
Pressure was applied to each plate to limit gas transfer between wells, and plates were incubated at –35 °C for 20 min.
Strips were removed from the plates and imaged with a flatbed scanner.
Images were converted to the CIELAB colour space and ROIs were defined around each blue reaction area, with each ROI representing a single sample.
ROIs were thresholded on the b* (0–128; ranging from blue to yellow) and L* (128–255; lightness) channels.
Blue intensity was calculated on a per pixel basis by subtracting the pixel L* value from 255.
Blue intensity values in a ROI were summed and the resulting value was normalized by the ROI area to quantify HCNp.
Dhurrin concentration was quantified using ultrahigh-performance liquid chromatography coupled with tandem mass spectrometry (UHPLC–MS/MS).
Leaf samples were harvested, dried at 60 °C for approximately 3 days and ground using a Bead Ruptor Elite tissue grinder (6 cycles at 4 m s –1 , 15 s each, with 10-s breaks).
A 100 mg aliquot of ground tissue was extracted with 750 μl of 50% methanol (v/v methanol and diH 2 O), incubated in a 75 °C water bath for 15 min, cooled to room temperature (10–15 min) and supplemented with an additional 750 μl of 50% methanol.
Samples were centrifuged at 11,000 rpm for 5 min.
A 30 μl aliquot of the supernatant was diluted with 270 μl of LC–MS-grade water (final 5% methanol, dilution factor 0.15 ml mg –1 ), transferred to a 2 ml glass HPLC vial and stored at –80 °C.
Quality control (QC) was maintained using pooled QC samples prepared from aliquots of individual extracts.
Blank extractions were also included.
Samples were stored at –80 °C until analysis.
One microlitre of extract was injected into a LX50 UHPLC system (PerkinElmer) equipped with a 20-µl sample loop (partial loop injection mode).
Chromatographic separation was performed using an Acquity UPLC HSS T3 column (1 × 50 mm, 1.8 µm; Waters) maintained at 45 °C.
The mobile phases consisted of water with 0.1% formic acid (A) and 100% acetonitrile (B).
The elution gradient started at 1% B for 0.5 min, ramped linearly to 99% B over 4.5 min, returned to 1% B at 5.2 min and was followed by a 2.8 min re-equilibration, for a total run time of 8 min.
The flow rate was set to 400 µl min –1 .
Detection was carried out on a PerkinElmer QSight 420 triple quadrupole mass spectrometer equipped with an electrospray ionization (ESI) source operating in positive mode and selected reaction monitoring (SRM).
The optimized SRM transitions for dhurrin were based on an authentic standard: Q1/Q3 = 333.9/144.9 for quantification and 333.9/184.9 for qualification.
Source parameters were as follows: drying gas temperature at 120 °C, hot-surface induced desolvation (HSID) temperature at 200 °C, electrospray voltage at 5,000 V and nebulizer gas flow at 350.
MS acquisition was scheduled around a retention time of 1.14 min with 1 min time windows.
The dwell time for each transition was 100 ms.
Data acquisition and processing were performed using Simplicity 3Q software (v.3.0, PerkinElmer).
Quantification was based on standard curves generated from authentic dhurrin standards using linear regression.
Concentrations are expressed as µg g –1 fresh weight, adjusted for sample weight and dilution factors.
For full details, see Supplementary Note 5 .
In brief, field-based measurements were collected in 2023 and 2024 across multiple locations captured key agronomic traits, including plant height, flowering time, panicle architecture and yield components, with data separated by replication and site to enable genotype × environment interaction analyses.
Phenotyping was conducted by the team at Donald Danforth Plant Science Center in 2023 and 2024.
High-resolution temporal vegetation indices, extracted from drone-based overhead imagery (Supplementary Table 4 ), further characterized canopy development and biomass dynamics over the growing season.
Controlled-environment phenotyping complemented field trials to quantify WUE and drought response under well-watered and water-limited conditions using automated indoor imaging systems.
Growth curve data across treatments enabled classification of genotypes by their drought response strategies, including early vigour and maintenance of growth under stress.
Greenhouse-based evaluations provided additional trait measurements under standardized conditions, including early vigour and developmental timing (Supplementary Data 12 ).
Visual documentation of phenotype diversity was collected across environments, including a curated set of representative field and greenhouse images for each genotype.
Environmental gradients associated with genomic variation in sorghum georeferenced lines
We used redundancy analysis (RDA) implemented with linear regression and principal component analysis to examine how environmental gradients explain genome-wide SNP variation among 443 sorghum landraces and to partition the variance in genomic diversity attributable to different sets of environmental variables.
RDA combines multiple regression with ordination to identify the environmental dimensions that best account for multilocus genetic structure and to quantify the proportion of total genomic variation explained by the environment.
Environmental variables included climatic factors, elevation, water balance, radiation, soil physicochemical properties and risk of striga.
All variables were standardized before analysis, and samples with missing data for any variable were excluded.
Genotype matrices were centred and regressed on the standardized environmental matrix; principal components of the fitted values defined the constrained axes (RDA axes) and their variance contributions.
The overall model fit, expressed as adjusted r 2 = 0.1948, indicated that the selected environmental variables jointly explained about 19.5% of genome-wide SNP variation.
Dhurrin and cyanide genome-wide association
GWAS were performed using a linear mixed model implemented in GEMMA (v.0.98.3) 115 .
Only directly called genotypes were used; no imputation was performed.
Variants were filtered to retain those with a MAF ≥ 0.05, resulting in 8963567 SNPs and short indels.
For each trait, association statistics were computed using the Wald test.
Markers with missing or undefined P values were excluded from downstream analyses.
The filtered results were used to generate Manhattan plots and to assess overlap with a priori candidate genes.
Significance was determined by FDR-correcting raw P values using the Benjamini–Hochberg method, implemented in the R internal function p.adjust.
In silico ligand docking of tyrosine to CYP79A1 protein variants
To evaluate the impact of sequence variants on the binding affinity of CYP79A1 for its substrate tyrosine, in silico ligand docking was performed using SwissDock ( https://www.swissdock.ch ).
Homology models in Protein Data Bank (PDB) format of wild-type and mutant CYP79A1 proteins were generated using SWISS-MODEL ( https://swissmodel.expasy.org ) using FASTA sequences as input and subsequently oriented in the endoplasmic reticulum (ER) membrane using the PPM3.0 server of the Orientation of Proteins in Membranes (OPM) database ( https://opm.phar.umich.edu/ppm_server ).
The resulting OPM-oriented PDB files were manually cleaned to remove water molecules and extraneous heteroatoms.
The MOL2 file for the substrate l -tyrosine was downloaded from the PubChem database and used as the ligand for docking.
PDB files of CYP79A1 containing either an alanine (A211) or valine (V211) at position 211, oriented in the membrane as described above, were used as the target structures.
Each docking run was performed under default settings, using the ‘Docking with AutoDock Vina’ option to scan for potential ligand-binding regions.
SwissDock returned a series of clusters, each associated with an estimated binding free energy (Δ G ) in kcal mol –1 .
The binding affinity of tyrosine to each CYP79A1 variant was compared by identifying the lowest predicted Δ G value across all clusters.
More negative Δ G values were interpreted as indicative of stronger substrate binding.
Predicted docking structures were visualized using PyMOL ( https://www.pymol.org/ ) to confirm that the substrate tyrosine localized in the active site of each protein model and to generate the renderings shown in Extended Data Fig.
5b,c .
Dhurrin BGC cis- element analysis
To test whether intergenic variants associated with seedling dhurrin content reside in regulatory regions with accessible chromatin and potential transcription factor (TF) binding sites, the Plant PAN 4.0 and SorghumBase 116 resources were used.
Genomic coordinates of intergenic variants in the BGC were examined using the Ensembl Plants genome browser hosted through SorghumBase to assess whether they overlapped with known accessible chromatin regions.
Specifically, the accessible chromatin region track derived from ATAC–seq profiling of 7-day-old S.
bicolor leaf tissue, as previously published 117 , was used, which provides a high-resolution map of chromatin accessibility across the genome.
To test whether these intergenic variants also overlapped with putative TF binding sites, each variant sequence was extracted along with 10 bp of flanking sequence on both sides.
These sequences were input into PlantPAN (v.4.0), a promoter and cis -element prediction tool that integrates binding site data from multiple plant-specific motif databases.
Only predicted TF-binding sites that directly overlapped the variant position were retained for downstream analysis, under the assumption that such overlap may indicate mutation of a functional cis -regulatory element.
To maximize confidence in predicted TF–DNA interactions, only hits that included both a known TF family and a corresponding Arabidopsis gene identifier were considered for downstream analysis.
Sequencing and genotyping of all Illumina libraries was performed once.
In cases when this resulted in duplicated genotypes, we avoided pseudo-replication by only retaining the library with higher depth of uniquely mapping reads for each duplicated genotype.
Phenotyping campaigns were spatially and temporally replicated as described in Supplementary Note 5 (with details regarding phenotype data collection and processing).
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Reference genome assembly and annotation files of all pangenome members are available online ( https://phytozome-next.jgi.doe.gov ).
Primary genome annotations and assemblies are available from EMBL/ENA with study and sample IDs reported in Supplementary Table 1 .
Pangenome graph (.gfa) and short-read variants called against the BTx623 V5 assembly (.vcf) are available on the Phytozome landing page for that genome.
Raw sequencing reads supporting genome assembly and annotation have been deposited into the SRA under the BioProject IDs listed in Supplementary Tables 2 and 6 .
Phenotype and sample metadata can be found in the corresponding supplementary data files, including SRA BioProjects for the short-read libraries (Supplementary Data 3 ).
Databases and datasets used in the study include Phytozome ( https://phytozome-next.jgi.doe.gov ), NCBI ( https://www.ncbi.nlm.nih.gov/ ) and OPM ( https://opm.phar.umich.edu/ ).
Source data are provided with this paper.
Richards, P.
Indigenous Agricultural Revolution: Ecology and Food Production in West Africa (Routledge, 2023).
Monk, R., Franks, C.
& Dahlberg, J.
in Yield Gains in Major U.S.
Field Crops (eds Smith, S.
et al.) 293–310 (American Society of Agronomy and Soil Science Society of America, 2015).
Rattunde, H.
F.
W.
et al.
Farmer participatory early-generation yield testing of sorghum in west Africa: possibilities to optimize genetic gains for yield in farmers’ fields.
Crop Sci.
56 , 2493–2505 (2016).
vom Brocke, K.
et al.
Participatory variety development for sorghum in Burkina Faso: farmers’ selection and farmers’ criteria.
Field Crops Res.
119 , 183–194 (2010).
Westengen, O.
T.
et al.
Ethnolinguistic structuring of sorghum genetic diversity in Africa and the role of local seed systems.
Proc.
Natl Acad.
Sci.
USA 111 , 14100–14105 (2014).
Article CAS PubMed PubMed Central ADS Google Scholar
Atlin, G.
N., Cairns, J.
E.
& Das, B.
Rapid breeding and varietal replacement are critical to adaptation of cropping systems in the developing world to climate change.
Glob.
Food Sec.
12 , 31–37 (2017).
Article PubMed PubMed Central Google Scholar
Cuevas, H.
E.
et al.
The evolution of photoperiod-insensitive flowering in sorghum.
A genomic model for panicoid grasses.
Mol.
Biol.
Evol.
33 , 2417–2428 (2016).
Article CAS PubMed PubMed Central Google Scholar
Abdulai, A.
L.
et al.
Latitude and date of sowing influences phenology of photoperiod-sensitive sorghums.
J.
Agron.
Crop Sci.
198 , 340–348 (2012).
Cobb, J.
N.
et al.
Enhancing the rate of genetic gain in public-sector plant breeding programs: lessons from the breeder’s equation.
Züchter Genet.
Breed.
Res.
132 , 627–645 (2019).
Walker, T.
S.
& Alwang, J.
Crop Improvement, Adoption and Impact of Improved Varieties in Food Crops in Sub-Saharan Africa (CABI Publishing, 2015).
Mauboussin, J.
C., Gueye, J.
& N’Diaye, M.
L’amélioration du sorgho au Sénégal.
Agron.
Trop.
32 , 303–310 (1977).
Muleta, K.
T.
et al.
The recent evolutionary rescue of a staple crop depended on over half a century of global germplasm exchange.
Sci.
Adv.
8 , eabj4633 (2022).
Kane, N.
A., Foncéka, D.
& Dalton, T.
J.
Crop Adaptation and Improvement for Drought-Prone Environments (New Prairie Press, 2022).
Woldeyohannes, A.
B.
et al.
Data-driven, participatory characterization of farmer varieties discloses teff breeding potential under current and future climates.
eLife https://doi.org/10.7554/eLife.80009 (2022).
Gesesse, C.
A.
et al.
Genomics-driven breeding for local adaptation of durum wheat is enhanced by farmers’ traditional knowledge.
Proc.
Natl Acad.
Sci.
USA 120 , e2205774119 (2023).
Paterson, A.
H.
et al.
The Sorghum bicolor genome and the diversification of grasses.
Nature 457 , 551–556 (2009).
Article CAS PubMed ADS Google Scholar
McCormick, R.
F.
et al.
The Sorghum bicolor reference genome: improved assembly, gene annotations, a transcriptome atlas, and signatures of genome organization.
Plant J.
93 , 338–354 (2018).
Mullet, J.
E.
High-biomass C4 grasses—filling the yield gap.
Plant Sci.
261 , 10–17 (2017).
Article CAS PubMed Google Scholar
Xin, Z.
et al.
Sorghum genetic, genomic, and breeding resources.
Planta 254 , 114 (2021).
Miller, J.
T.
et al.
Cloning and characterization of a centromere-specific repetitive DNA element from Sorghum bicolor .
Züchter Genet.
Breed.
Res.
96 , 832–839 (1998).
Laursen, T.
et al.
Characterization of a dynamic metabolon producing the defense compound dhurrin in sorghum.
Science 354 , 890–893 (2016).
Tao, Y.
et al.
Extensive variation within the pan-genome of cultivated and wild sorghum.
Nat.
Plants 7 , 766–773 (2021).
Zhang, Z.
et al.
Major impacts of widespread structural variation on sorghum.
Genome Res 34 , 286–299 (2024).
Jayakodi, M., Shim, H.
& Mascher, M.
What are we learning from plant pangenomes?
Annu.
Rev.
Plant Biol.
https://doi.org/10.1146/annurev-arplant-090823-015358 (2024).
Shang, L.
et al.
A super pan-genomic landscape of rice.
Cell Res.
32 , 878–896 (2022).
Cole, B.
et al.
Multi-season analysis reveals hundreds of drought-responsive genes in sorghum.
Plant J.
125 , e70657 (2026).
Liu, G.
& Godwin, I.
D.
Highly efficient sorghum transformation.
Plant Cell Rep.
31 , 999–1007 (2012).
Hufford, M.
B.
et al.
De novo assembly, annotation, and comparative analysis of 26 diverse maize genomes.
Science 373 , 655–662 (2021).
Bird, K.
A.
et al.
Structure and sequence evolution in the pennycress ( Thlaspi arvense ) pangenome.
Preprint at bioRxiv https://doi.org/10.1101/2025.09.26.678720 (2025).
Brůna, T., Sreedasyam, A., Harder, A.
M.
& Lovell, J.
T.
Evolutionary and methodological considerations when interpreting gene presence–absence variation in pangenomes.
NAR Genom.
Bioinform.
8 , lqag011 (2026).
Roeder, A.
H.
K.
et al.
Lost in translation: what we have learned from attributes that do not translate from Arabidopsis to other plants.
Plant Cell 37 , koaf036 (2025).
Gobena, D.
et al.
Mutation in sorghum LOW GERMINATION STIMULANT 1 alters strigolactones and causes Striga resistance.
Proc.
Natl Acad.
Sci.
USA 114 , 4471–4476 (2017).
Maina, F.
et al.
Delivering trait-enhanced varieties to African smallholders through a pangenomic breeding network.
Preprint at bioRxiv https://doi.org/10.1101/2025.08.07.667917 (2025).
Cavalet-Giorsa, E.
et al.
Origin and evolution of the bread wheat D genome.
Nature 633 , 848–855 (2024).
Healey, A.
L.
et al.
The complex polyploid genome architecture of sugarcane.
Nature 628 , 804–810 (2024).
Lin, Z.
et al.
Parallel domestication of the Shattering1 genes in cereals.
Nat.
Genet.
44 , 720–724 (2012).
Wu, X.
et al.
Genomic footprints of sorghum domestication and breeding selection for multiple end uses.
Mol.
Plant 15 , 537–551 (2022).
Fuller, D.
Q.
& Stevens, C.
J.
in Plants and People in the African Past (eds Mercuri, A.
M.
et al.) 427–452 (Springer, 2018).
Barnaud, A., Trigueros, G., McKey, D.
& Joly, H.
I.
High outcrossing rates in fields with mixed sorghum landraces: how are landraces maintained?.
Heredity 101 , 445–452 (2008).
Lasky, J.
R., Takou, M., Gamba, D.
& Keitt, T.
H.
Estimating scale-specific and localized spatial patterns in allele frequency.
Genetics 227 , iyae082 (2024).
Marcus, J., Ha, W., Barber, R.
F.
& Novembre, J.
Fast and flexible estimation of effective migration surfaces.
eLife 10 , e61927 (2021).
Petkova, D., Novembre, J.
& Stephens, M.
Visualizing spatial population structure with estimated effective migration surfaces.
Nat.
Genet.
48 , 94–100 (2016).
Wang, J., Hu, Z., Upadhyaya, H.
D.
& Morris, G.
P.
Genomic signatures of seed mass adaptation to global precipitation gradients in sorghum.
Heredity 124 , 108–121 (2020).
Szpiech, Z.
A., Novak, T.
E., Bailey, N.
P.
& Stevison, L.
S.
Application of a novel haplotype-based scan for local adaptation to study high-altitude adaptation in rhesus macaques.
Evol.
Lett.
5 , 408–421 (2021).
Ngara, R.
et al.
Identifying differentially expressed proteins in sorghum cell cultures exposed to osmotic stress.
Sci.
Rep.
8 , 8671 (2018).
Article PubMed PubMed Central ADS Google Scholar
Jeon, D., Kim, J.-B., Kang, B.-C.
& Kim, C.
Deciphering the genetic mechanisms of salt tolerance in Sorghum bicolor L.: key genes and SNP associations from comparative transcriptomic analyses.
Plants 12 , 2639 (2023).
Li, H.
et al.
A leucine-rich repeat-receptor-like kinase gene SbER2-1 from sorghum ( Sorghum bicolor L.) confers drought tolerance in maize.
BMC Genomics 20 , 737 (2019).
Krothapalli, K.
et al.
Forward genetics by genome sequencing reveals that rapid cyanide release deters insect herbivory of Sorghum bicolor .
Genetics 195 , 309–318 (2013).
Burke, J.
J.
et al.
Leaf dhurrin content is a quantitative measure of the level of pre- and postflowering drought tolerance in sorghum.
Crop Sci.
53 , 1056–1065 (2013).
Darbani, B.
et al.
The biosynthetic gene cluster for the cyanogenic glucoside dhurrin in Sorghum bicolor contains its co-expressed vacuolar MATE transporter.
Sci.
Rep.
6 , 37079 (2016).
Sibbesen, O., Koch, B., Halkier, B.
A.
& Møller, B.
L.
Isolation of the heme-thiolate enzyme cytochrome P-450TYR, which catalyzes the committed step in the biosynthesis of the cyanogenic glucoside dhurrin in Sorghum bicolo r (L.) Moench.
Proc.
Natl Acad.
Sci.
USA 91 , 9740–9744 (1994).
McCouch, S.
et al.
Agriculture: feeding the future.
Nature 499 , 23–24 (2013).
VanGessel, C.
J.
et al.
Ancient pangenomic origins of noncanonical NLR genes underlying the recent evolutionary rescue of a staple crop.
Sci.
Adv.
11 , eady1667 (2025).
Doyle, J.
J.
& Doyle, J.
L.
A rapid DNA isolation procedure for small quantities of fresh leaf tissue.
Phytochem.
Bull.
19 , 11–15 (1987).
Sreedasyam, A.
et al.
Genome resources for three modern cotton lines guide future breeding efforts.
Nat.
Plants 10 , 1039–1051 (2024).
Lovell, J.
et al.
Comparative analyses of four reference genomes reveal exceptional diversity and weak linked selection in the yellow monkeyflower ( Mimulus guttatus ) complex.
Mol.
Ecol.
Resour.
25 , e70012 (2025).
Koren, S.
et al.
Canu: scalable and accurate long-read assembly via adaptive k -mer weighting and repeat separation.
Genome Res.
27 , 722–736 (2017).
Xiao, C.-L.
et al.
MECAT: fast mapping, error correction, and de novo assembly for single-molecule sequencing reads.
Nat.
Methods 14 , 1072–1074 (2017).
Cheng, H., Concepcion, G.
T., Feng, X., Zhang, H.
& Li, H.
Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm.
Nat.
Methods 18 , 170–175 (2021).
Chin, C.-S.
et al.
Nonhybrid, finished microbial genome assemblies from long-read SMRT sequencing data.
Nat.
Methods 10 , 563–569 (2013).
Vaser, R., Sović, I., Nagarajan, N.
& Šikić, M.
Fast and accurate de novo genome assembly from long uncorrected reads.
Genome Res.
27 , 737–746 (2017).
Zhou, C.
et al.
Oatk: a de novo assembly tool for complex plant organelle genomes.
Preprint at bioRxiv https://doi.org/10.1101/2024.10.23.619857 (2024).
Li, H.
Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM.
Preprint at https://doi.org/10.48550/ARXIV.1303.3997 (2013).
McKenna, A.
et al.
The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data.
Genome Res.
20 , 1297–1303 (2010).
Goodstein, D.
M.
et al.
Phytozome: a comparative platform for green plant genomics.
Nucleic Acids Res.
40 , D1178–D1186 (2012).
Lovell, J.
T.
et al.
The genomic landscape of molecular responses to natural drought stress in Panicum hallii .
Nat.
Commun.
9 , 5213 (2018).
Wu, T.
D.
& Nacu, S.
Fast and SNP-tolerant detection of complex variants and splicing in short reads.
Bioinformatics 26 , 873–881 (2010).
Haas, B.
J.
et al.
Improving the Arabidopsis genome annotation using maximal transcript alignment assemblies.
Nucleic Acids Res.
31 , 5654–5666 (2003).
Smit, AFA, Hubley, R & Green, P.
RepeatMasker Open-4.0.
http://www.repeatmasker.org (2015).
Jones, P.
et al.
InterProScan 5: genome-scale protein function classification.
Bioinformatics 30 , 1236–1240 (2014).
Mistry, J.
et al.
Pfam: the protein families database in 2021.
Nucleic Acids Res.
49 , D412–D419 (2021).
Mi, H., Muruganujan, A., Ebert, D., Huang, X.
& Thomas, P.
D.
PANTHER version 14: more genomes, a new PANTHER GO-slim and improvements in enrichment analysis tools.
Nucleic Acids Res.
47 , D419–D426 (2018).
11.
Slater, G.
S.
C.
& Birney, E.
Automated generation of heuristics for biological sequence comparison.
BMC Bioinformatics 6 , 31 (2005).
Salamov, A.
A.
& Solovyev, V.
V.
Ab initio gene finding in Drosophila genomic DNA.
Genome Res.
10 , 516–522 (2000).
Stanke, M., Schöffmann, O., Morgenstern, B.
& Waack, S.
Gene prediction in eukaryotes with a generalized hidden Markov model that uses hints from external sources.
BMC Bioinformatics 7 , 62 (2006).
Hickey, G.
et al.
Pangenome graph construction from genome alignments with Minigraph-Cactus.
Nat.
Biotechnol.
42, 663–673 (2024).
Garrison, E.
et al.
Variation graph toolkit improves read mapping by representing genetic variation in the reference.
Nat.
Biotechnol.
36 , 875–879 (2018).
Guarracino, A., Heumos, S., Nahnsen, S., Prins, P.
& Garrison, E.
ODGI: understanding pangenome graphs.
Bioinformatics 38 , 3319–3326 (2022).
Beyer, W.
et al.
Sequence tube maps: making graph genomes intuitive to commuters.
Bioinformatics 35 , 5318–5320 (2019).
Garrison, E., Kronenberg, Z.
N., Dawson, E.
T., Pedersen, B.
S.
& Prins, P.
A spectrum of free software tools for processing the VCF variant call format: vcflib, bio-vcf, cyvcf2, hts-nim and slivar.
PLoS Comput.
Biol.
18 , e1009123 (2022).
Emms, D.
M.
& Kelly, S.
OrthoFinder: phylogenetic orthology inference for comparative genomics.
Genome Biol.
20 , 238 (2019).
Lovell, J.
T.
et al.
GENESPACE tracks regions of interest and gene copy number variation across multiple genomes.
eLife https://doi.org/10.7554/eLife.78526 (2022).
Goel, M., Sun, H., Jiao, W.-B.
& Schneeberger, K.
SyRI: finding genomic rearrangements and local sequence differences from whole-genome assemblies.
Genome Biol.
20 , 277 (2019).
Li, H.
Minimap2: pairwise alignment for nucleotide sequences.
Bioinformatics 34 , 3094–3100 (2018).
Parmigiani, L., Garrison, E., Stoye, J., Marschall, T.
& Doerr, D.
Panacus: fast and exact pangenome growth and core size estimation.
Bioinformatics https://doi.org/10.1093/bioinformatics/btae720 (2024).
Li, H.
et al.
The Sequence Alignment/Map format and SAMtools.
Bioinformatics 25 , 2078–2079 (2009).
Koboldt, D.
C.
et al.
VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing.
Genome Res.
22 , 568–576 (2012).
Delaneau, O., Marchini, J.
& Zagury, J.-F.
A linear complexity phasing method for thousands of genomes.
Nat.
Methods 9 , 179–181 (2011).
Voichek, Y.
& Weigel, D.
Identifying genetic variants underlying phenotypic variation in plants without complete genomes.
Nat.
Genet.
52 , 534–540 (2020).
Spindel, J.
E.
et al.
Association mapping by aerial drone reveals 213 genetic associations for Sorghum bicolor biomass traits under drought.
BMC Genomics 19 , 679 (2018).
Brenton, Z.
W.
et al.
A genomic resource for the development, improvement, and exploitation of sorghum for bioenergy.
Genetics 204 , 21–33 (2016).
Casa, A.
M.
et al.
Community resources and strategies for association mapping in sorghum.
Crop Sci.
48 , 30–40 (2008).
Lozano, R.
et al.
Comparative evolutionary genetics of deleterious load in sorghum and maize.
Nat.
Plants 7 , 17–24 (2021).
Wang, T., Crawford, M.
M.
& Tuinstra, M.
R.
A novel transfer learning framework for sorghum biomass prediction using UAV-based remote sensing data and genetic markers.
Front.
Plant Sci.
14 , 1138479 (2023).
Masjedi, A., Crawford, M.
M., Carpenter, N.
R.
& Tuinstra, M.
R.
Multi-temporal predictive modelling of sorghum biomass using UAV-based hyperspectral and LiDAR data.
Remote Sens.
12 , 3587 (2020).
Billot, C.
et al.
Massive sorghum collection genotyped with SSR markers to enhance use of global genetic resources.
PLoS ONE 8 , e59714 (2013).
Garin, V.
et al.
Characterization of adaptation mechanisms in sorghum using a multireference back-cross nested association mapping design and envirotyping.
Genetics https://doi.org/10.1093/genetics/iyae003 (2024).
Faye, J.
M.
et al.
Genomic signatures of adaptation to Sahelian and Soudanian climates in sorghum landraces of Senegal.
Ecol.
Evol.
9 , 6038–6051 (2019).
Alexander, D.
H., Novembre, J.
& Lange, K.
Fast model-based estimation of ancestry in unrelated individuals.
Genome Res.
19 , 1655–1664 (2009).
Purcell, S.
et al.
PLINK: a tool set for whole-genome association and population-based linkage analyses.
Am.
J.
Hum.
Genet.
81 , 559–575 (2007).
Li, H.
A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data.
Bioinformatics 27 , 2987–2993 (2011).
Browning, B.
L., Tian, X., Zhou, Y.
& Browning, S.
R.
Fast two-stage phasing of large-scale sequence data.
Am.
J.
Hum.
Genet.
108 , 1880–1890 (2021).
Cingolani, P.
et al.
A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3.
Fly 6 , 80–92 (2012).
Rhoné, B.
et al.
Pearl millet genomic vulnerability to climate change in West Africa highlights the need for regional collaboration.
Nat.
Commun.
11 , 5274 (2020).
Lasky, J.
R., Takou, M., Gamba, D.
& Keitt, T.
H.
Estimating scale-specific and localized spatial patterns in allele frequency.
Genetics https://doi.org/10.1101/2022.03.21.485229 (2024).
M McLaughlin, C.
et al.
Maladaptation in cereal crop landraces following a soot-producing climate catastrophe.
Nat.
Commun.
16 , 4289 (2025).
Kemanian, A.
R.
et al.
The Cycles agroecosystem model: fundamentals, testing, and applications.
Comput.
Electron.
Agric.
227 , 109510 (2024).
Shi, Y., Montes, F.
& Kemanian, A.
R.
Cycles-L: a coupled, 3-D, land surface, hydrologic, and agroecosystem landscape model.
Water Resour.
Res .
https://doi.org/10.1029/2022WR033453 (2023).
Rodell, M.
et al.
The global land data assimilation system.
Bull.
Am.
Meteorol.
Soc.
85 , 381–394 (2004).
Hengl, T.
et al.
SoilGrids250m: global gridded soil information based on machine learning.
PLoS ONE 12 , e0169748 (2017).
Leonard, L.
& Duffy, C.
Visualization workflows for level-12 HUC scales: towards an expert system for watershed analysis in a distributed computing environment.
Environ.
Model.
Softw.
78 , 163–178 (2016).
Leonard, L.
& Duffy, C.
J.
Automating data-model workflows at a level 12 HUC scale: watershed modeling in a distributed computing environment.
Environ.
Model.
Softw.
61 , 174–190 (2014).
Leonard, L.
& Duffy, C.
J.
Essential Terrestrial Variable data workflows for distributed water resources modeling.
Environ.
Model.
Softw.
50 , 85–96 (2013).
Fick, S.
E.
& Hijmans, R.
J.
WorldClim 2: new 1km spatial resolution climate surfaces for global land areas.
Int.
J.
Climatol.
37 , 4302–4315 (2017).
Zhou, X.
& Stephens, M.
Genome-wide efficient mixed-model analysis for association studies.
Nat.
Genet.
44 , 821–824 (2012).
Gladman, N.
et al.
SorghumBase: a web-based portal for sorghum genetic information and community advancement.
Planta 255 , 35 (2022).
Lu, Z.
et al.
The prevalence, evolution and chromatin signatures of plant regulatory elements.
Nat.
Plants 5 , 1250–1259 (2019).
The work (proposals: 503014, 504730, 2015; Award DOIs: 10.46936/10.25585/60001093, 10.46936/10.25585/60001224, 10.46936/10.25585/60001015) conducted by the US DOE Joint Genome Institute ( https://ror.org/04xm1d337 ), a DOE Office of Science User Facility, is supported by the Office of Science of the US DOE operated under contract number DE-AC02-05CH11231.
This work was also supported by Gates Foundation projects: ‘Green Evolution—Accelerating Dryland Cereals Improvement for Africa’ (INV-053669), the ‘Sorghum Genomics Toolbox: TERRA Partnership (OPP1129603)’, and ‘Mining useful alleles for climate change adaptation from CGIAR gene banks’.
The information presented herein was funded in part by the Advanced Research Projects Agency-Energy (ARPA-E), US Department of Energy, under Award Numbers DE-AR0000594.
S.M.B., D.K.
and T.T.
acknowledge support from NIOO via grant OPP1082853 ‘RSM Systems Biology for Sorghum’.
J.T.L.
was supported by the Center for Bioenergy Innovation (US DOE, Office of Science, Biological and Environmental Research under contract number ERKP886).
P.G.L.
was supported by the US Cooperative Extension Service through the Division of Agriculture and Natural Resources of the University of California and DOE Grant DE-SC0014081.
J.E.M.
was supported by the Great Lakes Bioenergy Research Center (DOE BER Office of Science Grant/Award: DE-SC0018409).
M.E.H.
is supported by the DOE Center for Advanced Bioenergy and Bioproducts Innovation (US DOE, Office of Science, Biological and Environmental Research Program under Award Number DE-SC0018420).
This study is made possible by the support of the American People provided to the Feed the Future Innovation Lab for Collaborative Research on Sorghum and Millet through the United States Agency for International Development (USAID) under associate award no.
AID-OAA-A13-00047, ‘Feed the Future Innovation Lab for Genomics-Assisted Sorghum Breeding’ and ‘Feed the Future Innovation Lab for Crop Improvement’.
The contents are the sole responsibility of the authors and do not necessarily reflect the views of USAID or the US Government.
We dedicate this work to the memory of Todd C.
Mockler, a founding contributor to the sorghum pangenome effort.
We honour his vision by advancing this work and building on the groundwork he helped establish.
His loss is felt deeply, and his legacy lives on through the data, tools and collaborations he inspired.
Present address: Experimental and Computational Plant Development and Plant Stress Resilience, Institute of Environmental Biology, Department of Biology, Utrecht University, Utrecht, The Netherlands
These authors contributed equally: Geoffrey P.
Morris, Avril M.
Harder, Adam L.
Healey, Chloee M.
McLaughlin, Joanna L.
Rifkin
Department of Soil and Crop Sciences, Colorado State University, Fort Collins, CO, USA
Geoffrey P.
Morris, Clara Cruet-Burgos, John J.
Spiekerman, Carl J.
VanGessel, Kristen K.
Johnson, John K.
McKay & Brian R.
Rice
Genome Sequencing Center, HudsonAlpha Institute for Biotechnology, Huntsville, AL, USA
Avril M.
Harder, Adam L.
Healey, Chloee M.
McLaughlin, Joanna L.
Rifkin, Jerry W.
Jenkins, Lori Beth Boston, Dave Flowers, Jane Grimwood, Sujan Mamidi, Christopher Plott, Avinash Sreedasyam, Rachel Walstead, Jenell Webber, Melissa Williams, Jeremy Schmutz & John T.
Lovell
Department of Horticulture and Landscape Architecture, Colorado State University, Fort Collins, CO, USA
Chloee M.
McLaughlin, Jacqueline M.
Chaparro & Jessica E.
Prenni
US Department of Energy Joint Genome Institute, Lawrence Berkeley National Laboratory, Berkeley, CA, USA
Joanna L.
Rifkin, Shengqiang Shu, Kerrie Barry, David Goodstein, Avinash Sreedasyam, John P.
Vogel, Jeremy Schmutz & John T.
Lovell
Donald Danforth Plant Science Center, St Louis, MO, USA
Erica Agnew, Ivan Baxter, Nathaniel Eck, Andrea L.
Eveland, Boubacar Gano, Marie de Gracia Coquerel, Scott Lee, Philip Ozersky, Jocelyn Saxton, Todd C.
Mockler & Nadia Shakoor
CIRAD, UMR AGAP, University Montpellier and Institut Agro, Montpellier, France
Alain Audebert, Gregory Beurier, Daniel Fonceka, Delphine Luquet & Jean-François Rami
Department of Plant and Environmental Sciences, Clemson University, Clemson, SC, USA
Richard E.
Boyles, Chaney Courtney, Stephen Kresovich & Trevor W.
Rife
Howard Hughes Medical Institute, University of California, Davis, Davis, CA, USA
Department of Plant Biology and Genome Center, University of California, Davis, Davis, CA, USA
Arizona Genomics Institute, University of Arizona, Tucson, AZ, USA
Victoria Bunting & Jayson Talag
Faculté de Pharmacie, Université des Sciences des Techniques et des Technologies (USTTB), Bamako, Mali
International Crops Research Institute for the Semi-Arid Tropics (ICRISAT), Patancheru, India
Santosh Deshpande & Jana Kholova
Institut Sénégalais de Recherches Agricoles, Thiès, Sénégal
Cyril Diatta, Jacques M.
Faye & Bassirou Sine
DOE Center for Advanced Bioenergy and Bioproducts Innovation (CABBI), University of Illinois at Urbana-Champaign, Urbana, IL, USA
Department of Crop Sciences, University of Illinois at Urbana-Champaign, Urbana, IL, USA
Center for Craniofacial Molecular Biology, Herman Ostrow School of Dentistry, University of Southern California, Los Angeles, CA, USA
Institut d’Economie Rurale du Mali, Bamako, Mali
Innovative Genomics Institute, University of California Berkeley, Berkeley, CA, USA
Department of Plant and Microbial Biology, University of California Berkeley, Berkeley, CA, USA
Lone Wolf Biotech, St Louis, MO, USA
Institut National de la Recherche Agronomique du Niger, Niamey, Niger
The Plant Molecular and Cellular Biology Laboratory, The Salk Institute for Biological Studies, La Jolla, CA, USA
Department of Cell and Developmental Biology, School of Biological Sciences, University of California, San Diego, La Jolla, CA, USA
Department of Science and Conservation, San Diego Botanical Garden, Encinitas, CA, USA
Center for Marine Biotechnology and Biomedicine, University of California, San Diego, La Jolla, CA, USA
Ethiopian Institute of Agricultural Research, Addis Ababa, Ethiopia
Texas A&M University, College Station, TX, USA
CHIBAS, Centre Haïtien d’Innovation en Biotechnologies et pour une Agriculture Soutenable, Croix des Bouquets, Haïti
Department of Agronomy, Purdue University, West Lafayette, IN, USA
IRD, DIADE Unit, Université de Montpellier, Montpellier, France
School of Tropical Agriculture and Forestry, Hainan University, Haikou, China
Department of Biology, Pennsylvania State University, University Park, PA, USA
Department of Agronomy and Horticulture, University of Nebraska, Lincoln, NE, USA
Search author on: PubMed Google Scholar
All authors contributed significantly over the 15 years of this project.
The following summarizes contributions as they relate to this article.
Writing: J.T.L., C.M.M., J.L.R., G.P.M., N.S., J.R.L., J.K.M., J.
Schmutz and B.R.R.
Sequencing, data hosting and sample preparation: J.G., J.
Schmutz, L.B.B., J.T., D.G., M.W., J.
Webber and R.L.
Genome assembly and annotation: J.W.J., C.P., S.S., R.W., J.T.L.
and A.S.
Variant detection: S.M., A.L.H., D.
Flowers, C.M.M.
and J.L.R.
Pangenomics: A.M.H., A.L.H., J.L.R., A.S., C.M.M.
and J.T.L.
Analyses of population and landscape genetics: J.L.R., C.M.M., Y.X., J.T.L., J.
Wang and C.J.V.
Genetic resources: J.P.V., J.M., P.L., S.K., A.L.E., M.E.H., G.P.M., G.P., M.R.T., S.D., N.T., C.D., J.M.F., D.K., F.M.
and T.T.M.
Field work, phenotyping and quantitative genetics: N.S., P.O., C.J.V., J.
Schmutz, B.R.R., N.E., T.W.R., C.C., M.d.G.C., R.E.B., S.L., J.
Saxton, B.G., K.J., J.M.C., J.E.P.
and K.K.J.
Project and sample management: K.B., J.
Schmutz, J.G., G.P.M., J.T.L., C.C.-B., I.B., D.G., E.A., N.S., V.B.
and J.S.B.D.
Conceptualization and funding: G.P.M., J.T.L., J.
Schmutz, T.P.M., N.S., J.V., J.L.R., T.P.M., B.R.R., C.C.-B., I.B., J.K.M., S.M.B., G.P., A.A., G.B., J.K., J.-F.R., B.S., N.T., V.V., D.L., M.K., D.
Foceka and T.P.M.
Correspondence to Geoffrey P.
Morris , Nadia Shakoor or John T.
Lovell .
The authors declare no competing interests.
Nature thanks Damaris Odeny, Scott Sattler and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig.
1 Genetic principal components.
Coordinates from the diversity panel of 1,984 unique genotypes (short read-based) principal components are shown for the first two axes (top) and first and third axes (bottom), color-coded by the ten admixture subpopulations (left) and the five major botanical types (right).
The 33 members of the pangenome reference are labeled with the colors of the bounding boxes matching the colors of groupings in that plot.
Extended Data Fig.
2 Whole-plant phenotyping of the reference pangenome and diversity panel members.
a Distribution of breeding values of all genotypes grown in our two field sites across two years, phenotyped manually for three key traits ( n genotypes for days to anthesis = 237, n plant height = 242, n biomass = 242).
The whole panel distribution is visualized as a histogram with 50 bins per panel, and the positions of pangenome members are flagged in the rug below the histogram.
b Stomatal conductance ( g s ) phenotyped on a LICOR infrared gas analyzer plotted in the same format as panel A ( n = 241).
c Whole-plant water use-efficiency across 30 of the pangenome reference members, defined as the amount of water used per total plant area gained (phenotyped via high-throughput imaging).
d Manually collected root biomass accumulation under well-watered (100% of field capacity, FC) and water-limited (40% of FC) treatments.
Genotype means for 26 of the pangenome reference members are plotted overlaying the genetic correlation ± 95% confidence intervals (Pearsons’ r = 0.40, P < 0.001).
e Field collected growth curves defined by two UAV-derived vegetation indices: green Normalized Difference Vegetation Index (NDVI; (NIR mean - green mean)/(NIR mean + green mean)), and Modified Chlorophyll Absorption in Reflectance Index (MCARI; (red edge mean - red mean - 0.2 * (red edge mean - green mean)) * (red edge mean/red mean), plotted from 25 to 70 days after sowing for 430 genotypes.
The diversity panel members are thin grey lines, the pangenome reference members are thicker lines.
Four references that span the diversity of measurements are colored and labeled.
Extended Data Fig.
3 Details of climate- and spatial-explicit population genetics.
a Wavelet dissimilarity plots of georeferenced samples were divided into six groups with partitioning around medoid (‘PAM’) clustering and split into drought-associated and non-drought-associated climates within each PAM cluster.
b The geographic distribution of georeferenced diversity panel members, colored by PAM clusters and drought status.
c Untransformed values of wavelet dissimilarity in the three species presented in Fig.
4a .
d Environmental gradients associated with genomic differentiation are plotted for the first two constrained axes and overlaid biplot vectors for representative environmental variables.
Points are n = 421 georeferenced diversity panel resequencing libraries, symbol-coded by botanical type, and colors follow sh1 haplotypes as in Fig.
3 .
Arrow direction and length represent the orientation and relative strength of environmental associations in the RDA space.
e The 1,984 resequencing libraries were split by botanical type (left: five ‘pure’ botanical types [‘Caud’ = Caudatum], intermediate [‘IM’], and unknown [NA]), and k = 10 genetic subpopulations (right).
In each grouping the proportion of libraries with strong (>75% identity - lighter colors), and weaker (33-75% identity - darker colors) are plotted for each sh1 haplotype.
The total number of libraries in each grouping is printed above the plot.
Bars are ordered by decreasing assignment to sh1 -1.
f Summary gene ontology (GO) enrichments showing the significance (x-axis) and contribution of genes (numbers next to points) for the top 10 enrichments for wavelet dissimilarity scans at geographic scales of 790 km and 1,119 km.
Extended Data Fig.
4 Extended haplotype block positions.
Extended haplotype blocks, indicative of selective sweeps, plotted overall (global) and for the 5 geographic regions with sufficient replication of both drought-prone (light red background) and non-drought prone (light blue background) geographic regions.
The 8,109 visualized blocks are simplified from the 30,289 raw blocks so that small intervals are visible by adding 25 kb buffers to all significant blocks and reducing overlapping intervals to a single range.
Plot colors follow that of the map in Extended Data Fig.
3a,b .
Extended Data Fig.
5 Detailed exploration of the dhurrin BGC.
a GWAS ‘Manhattan’ plots for dhurrin concentration, and hydrogen cyanide potential at the early seedling (3-4 leaf) and early vegetative (7-8 leaf) growth stages across the BTx623 V5 genome coordinate system.
Dhurrin concentration association plot is the same as in Fig.
5a .
Wald test P -values (-log 10 scale) are reported along with the FDR (alpha = 0.05) corrected threshold for genome-wide significance.
b Homology models of the CYP79A1 enzyme containing either an alanine (left, A211) or valine (right, V211) residue at position 211 were generated in SWISS-MODEL and oriented within the ER membrane using the PPM3.0 server of the Orientation of Proteins in Membranes (OPM) database ( https://opm.phar.umich.edu/ppm_server ).
In each model, residue 211 is outlined in green.
c Predicted docking poses of the substrate L-tyrosine in the binding pocket of CYP79A1 A211 (left) and V211 (right) variants.
Ligand docking was performed using SwissDock with the AutoDock Vina protocol.
The most energetically favorable pose for each variant is shown, along with the predicted binding free energy (ΔG, kcal/mol), where more negative values indicate stronger binding affinity.
d Phenotypic effects of resequenced short variant allelic combinations across six relatively unlinked sites in the BGC.
Here, we include two additional SNPs that show high levels of missingness (SNP3-4: Chr01: 1,094,821, 1,149,356).
INDEL1-2 and SNP1-2 are the same as in Fig.
5c .
BTx623 genotype homozygote is yellow, heterozygote is green, alternative homozygote is blue, and missing genotype call is gray.
The number of phenotyped samples with a given allelic combination is denoted to the left (unlabeled rows have only a single library with that combination of genotypes).
e The log-transformed seedling dhurrin content of plants corresponding to allelic combinations.
Supplementary Information (download PDF )
Supplementary Notes 1–5, Supplementary Figs.
1–3 and Supplementary Tables 1–4.
Reporting Summary (download PDF )
Supplementary Data (download ZIP )
Peer Review file (download PDF )
Source Data Figs.
2, 4, 5 and Extended Data Figs.
2–5 (download XLSX )
Version of record : 11 March 2026
DOI : https://doi.org/10.1038/s41586-026-10229-9
Related Stories
Source: This article was originally published by Nature News
Read Full Original Article →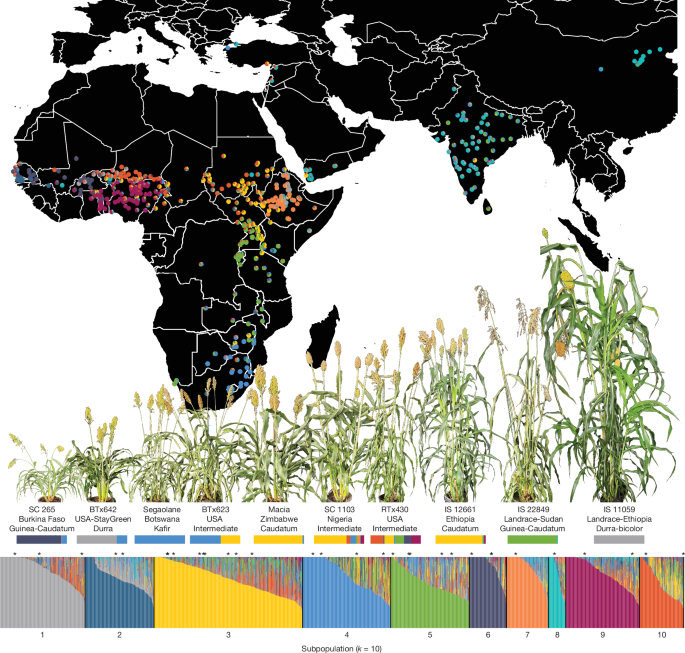



Comments (0)
No comments yet. Be the first to comment!
Leave a Comment