Nature ( 2026 ) Cite this article
Despite promising data showing that circulating tumour DNA (ctDNA) dynamics during treatment can inform real-time tumour response and recurrence risk 1 , how best to translate these insights into actionable clinical decision-making remains unclear.
Here we report results from the EP-STAR trial—a multi-centre, ctDNA-driven, risk-adapted, non-randomized phase II study ( NCT04072107 ; ClinicalTrials.gov) testing whether a risk-adaptive treatment (RAT) strategy guided by on-treatment ctDNA dynamics can meaningfully improve survival, using nasopharyngeal carcinoma as a model.
Eligible patients were enrolled and began treatment with standard-of-care gemcitabine–cisplatin neoadjuvant chemotherapy (GP-NAC; the P in this abbreviation stands for platinum) 2 , followed by RAT or standard-of-care chemoradiotherapy guided by ctDNA clearance trajectory during GP-NAC.
Protocol-eligible patients who did not receive RAT, drawn from a prospectively registered ctDNA biomarker cohort ( NCT03855020 ) 3 , served as a non-randomized, contemporaneous no-RAT external cohort.
The primary end-point was failure-free survival (FFS) in the RAT group.
After a median follow-up of 47.3 months, the 3-year FFS was 89.1% (83.2–95.0%) in the RAT group ( n = 110).
Patients who received RAT showed significantly improved FFS ( P = 0.003, log–rank test) compared with the no-RAT external cohort (hazard ratio = 0.41 [0.23–0.75]; P = 0.004, Cox regression model).
The RAT strategy was well-tolerated with no treatment-related deaths.
Collectively, these data show that a ctDNA-driven RAT paradigm could be a promising strategy to improve survival, challenging the conventional fixed-course, static treatment approach.
This is a preview of subscription content, access via your institution
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
Receive 51 print issues and online access
Prices may be subject to local taxes which are calculated during checkout
Fig.
2: Flow chart of participant enrolment and treatment.
Fig.
3: Failure-free survival of patients in the ITT population.
Fig.
4: Survival outcomes of the RAT group versus the no-RAT external cohort.
Similar content being viewed by others
Circulating tumor DNA-guided adjuvant therapy in locally advanced colon cancer: the randomized phase 2/3 DYNAMIC-III trial
Improved overall survival is associated with adjuvant chemotherapy after definitive concurrent chemoradiotherapy for N3 nasopharyngeal cancer
ctDNA guiding adjuvant immunotherapy in urothelial carcinoma
De-identified, anonymized, participant-level data have been deposited in the Mendeley Data repository ( https://doi.org/10.17632/wk275rgy98.1 ) and are publicly available as of the date of publication.
The scRNA-seq sequencing data were obtained from published research 5 , and downloaded from the CNGB Sequence Archive ( https://db.cngb.org/cnsa/ ) under the accession number CNP0001341 .
The public database MSigDB, which provides a resource of annotated gene sets for use, is available at https://www.gsea-msigdb.org .
Source data are provided with this paper.
Ignatiadis, M., Sledge, G.
W.
& Jeffrey, S.
S.
Liquid biopsy enters the clinic—implementation issues and future challenges.
Nat.
Rev.
Clin.
Oncol.
18 , 297–312 (2021).
Zhang, Y.
et al.
Gemcitabine and cisplatin induction chemotherapy in nasopharyngeal carcinoma.
N.
Engl.
J.
Med.
381 , 1124–1135 (2019).
Article CAS PubMed Google Scholar
Lv, J.
et al.
Longitudinal on-treatment circulating tumor DNA as a biomarker for real-time dynamic risk monitoring in cancer patients: the EP-SEASON study.
Cancer Cell 42 , 1401–1414 (2024).
Kurtz, D.
M.
et al.
Dynamic risk profiling using serial tumor biomarkers for personalized outcome prediction.
Cell 178 , 699–713 (2019).
Article CAS PubMed PubMed Central Google Scholar
Lv, J.
et al.
The tumor immune microenvironment of nasopharyngeal carcinoma after gemcitabine plus cisplatin treatment.
Nat.
Med.
29 , 1424–1436 (2023).
Wang, X.
Q.
et al.
Spatial predictors of immunotherapy response in triple-negative breast cancer.
Nature 621 , 868–876 (2023).
Article ADS CAS PubMed PubMed Central Google Scholar
Cohen, S.
A., Liu, M.
C.
& Aleshin, A.
Practical recommendations for using ctDNA in clinical decision making.
Nature 619 , 259–268 (2023).
Article ADS CAS PubMed Google Scholar
Nabet, B.
Y.
et al.
Noninvasive early identification of therapeutic benefit from immune checkpoint inhibition.
Cell 183 , 363–376 (2020).
Pan, Y.
et al.
Dynamic circulating tumor DNA during chemoradiotherapy predicts clinical outcomes for locally advanced non-small cell lung cancer patients.
Cancer Cell 41 , 1763–1773 (2023).
Lv, J.
et al.
Liquid biopsy tracking during sequential chemo-radiotherapy identifies distinct prognostic phenotypes in nasopharyngeal carcinoma.
Nat.
Commun.
10 , 3941 (2019).
Article ADS PubMed PubMed Central Google Scholar
Lv, J.
et al.
Improving on-treatment risk stratification of cancer patients with refined response classification and integration of circulating tumor DNA kinetics.
BMC Med 20 , 268 (2022).
Brierley, J.
D.
et al.
(eds) TNM Classification of Malignant Tumours 8th edn (Wiley-Blackwell, 2016).
Chen, Y.
P.
et al.
Low-dose metronomic chemotherapy improves tumor control in nasopharyngeal carcinoma.
Cancer Commun.
42 , 909–912 (2022).
Liu, X.
et al.
Induction-concurrent chemoradiotherapy with or without sintilimab in patients with locoregionally advanced nasopharyngeal carcinoma in China (CONTINUUM): a multicentre, open-label, parallel-group, randomised, controlled, phase 3 trial.
Lancet 403 , 2720–2731 (2024).
Liang, Y.-L.
et al.
Adjuvant PD-1 blockade with camrelizumab for nasopharyngeal carcinoma: the DIPPER randomized clinical trial.
J.
Am.
Med.
Assoc.
333 , 1589–1598 (2025).
Dong, S.
et al.
Circulating tumor DNA-guided de-escalation targeted therapy for advanced non-small cell lung cancer: a nonrandomized controlled trial.
JAMA Oncol.
10 , 932–940 (2024).
Article PubMed PubMed Central Google Scholar
Tie, J.
et al.
Circulating tumor DNA analysis guiding adjuvant therapy in stage II colon cancer.
N.
Engl.
J.
Med.
386 , 2261–2272 (2022).
Tie, J.
et al.
Circulating tumor DNA analysis guiding adjuvant therapy in stage II colon cancer: 5-year outcomes of the randomized DYNAMIC trial.
Nat.
Med.
31 , 1509–1518 (2025).
Nakamura, Y.
et al.
ctDNA-based molecular residual disease and survival in resectable colorectal cancer.
Nat.
Med.
30 , 3272–3283 (2024).
Tie, J.
et al.
Circulating tumor DNA-guided adjuvant therapy in locally advanced colon cancer: the randomized phase 2/3 DYNAMIC-III trial.
Nat.
Med.
31 , 4291–4300 (2025).
Chen, L.
et al.
Postoperative human papilloma virus circulating tumor DNA guided adjuvant therapy for human papilloma virus-related oropharyngeal carcinoma (PATH study).
Int.
J.
Radiat.
Oncol.
Biol.
Phys.
124 , 676–685 (2026).
Huang, Z.
et al.
Plasma Epstein–Barr virus DNA temporal clearance pattern during induction-concurrent (chemo)radiation therapy for risk stratification in nasopharyngeal carcinoma.
Int.
J.
Radiat.
Oncol.
Biol.
Phys.
123 , 129–140 (2025).
Chen, Y.
P.
et al.
Metronomic capecitabine as adjuvant therapy in locoregionally advanced nasopharyngeal carcinoma: a multicentre, open-label, parallel-group, randomised, controlled, phase 3 trial.
Lancet 398 , 303–313 (2021).
Tang, L.-L.
et al.
CACA guidelines for holistic integrative management of nasopharyngeal carcinoma.
Holist.
Integr.
Oncol.
2 , 24, (2023).
Liu, S.
L.
et al.
Neoadjuvant and adjuvant toripalimab for locoregionally advanced nasopharyngeal carcinoma: a randomised, single-centre, double-blind, placebo-controlled, phase 2 trial.
Lancet Oncol.
25 , 1563–1575 (2024).
Langer, C.
J.
et al.
Carboplatin and pemetrexed with or without pembrolizumab for advanced, non-squamous non-small-cell lung cancer: a randomised, phase 2 cohort of the open-label KEYNOTE-021 study.
Lancet Oncol.
17 , 1497–1508 (2016).
Rha, S.
Y.
et al.
Pembrolizumab plus chemotherapy versus placebo plus chemotherapy for HER2-negative advanced gastric cancer (KEYNOTE-859): a multicentre, randomised, double-blind, phase 3 trial.
Lancet Oncol.
24 , 1181–1195 (2023).
Le, Q.
T.
et al.
An international collaboration to harmonize the quantitative plasma Epstein–Barr virus DNA assay for future biomarker-guided trials in nasopharyngeal carcinoma.
Clin.
Cancer Res.
19 , 2208–2215 (2013).
Tang, S.
Q.
et al.
Identifying optimal clinical trial candidates for locoregionally advanced nasopharyngeal carcinoma: analysis of 9468 real-world cases and validation by two phase 3 multicentre, randomised controlled trial.
Radiother.
Oncol.
167 , 179–186 (2022).
Lin, J.
C.
et al.
Quantification of plasma Epstein–Barr virus DNA in patients with advanced nasopharyngeal carcinoma.
N.
Engl.
J.
Med.
350 , 2461–2470 (2004).
Chan, K.
C.
A.
et al.
Analysis of plasma Epstein–Barr virus DNA to screen for nasopharyngeal cancer.
N.
Engl.
J.
Med.
377 , 513–522 (2017).
Lo, Y.
M.
et al.
Quantitative analysis of cell-free Epstein–Barr virus DNA in plasma of patients with nasopharyngeal carcinoma.
Cancer Res.
59 , 1188–1191 (1999).
Schmidt, R.
et al.
Sample size calculation for the one-sample log-rank test.
Stat.
Med.
34 , 1031–1040 (2015).
Article MathSciNet PubMed Google Scholar
Borgan, Ø & Liestøl, K.
A note on confidence intervals and bands for the survival function based on transformations.
Scand.
J.
Stat.
17 , 35–41 (1990).
Schoenfeld, D.
A.
Sample-size formula for the proportional-hazards regression model.
Biometrics 39 , 499–503 (1983).
Kurz, C.
F., Krzywinski, M.
& Altman, N.
Propensity score matching.
Nat.
Methods 21 , 1770–1772 (2024).
Osoba, D., Rodrigues, G., Myles, J., Zee, B.
& Pater, J.
Interpreting the significance of changes in health-related quality-of-life scores.
J.
Clin.
Oncol.
16 , 139–144 (1998).
Bertram, M.
Y.
et al.
Cost-effectiveness thresholds: pros and cons.
Bull.
World Health Organ.
94 , 925–930 (2016).
This trial was supported by grants from the Science Fund for Creative Research Groups of the National Natural Science Foundation of China (82521003 to Y.S.), the National Natural Science Foundation of China for Excellent Young Scientists Fund (82522065 to J.L.), the National Natural Science Foundation of China (82441026 to Y.S., 92259202 to Y.S.
and 81803105 to J.-B.L.), the Fundamental and Interdisciplinary Disciplines Breakthrough Plan of the Ministry of Education of China (JYB2025XDXM611 to J.M.), the Guangdong Basic and Applied Basic Research Foundation (2024A1515030248 to G.-Q.Z.), the Young Talents Program of SYSUCC (YTP-SYSUCC-0103 to J.L.), the Cancer Innovative Research Program of SYSUCC (CIRP-SYSUCC-0010 to Y.S.), the Overseas Expertise Introduction Project for Discipline Innovation (111 Project) (B14035 to J.M.) and the National Medical Research Council Singapore Clinician Scientist Award (NMRC/CSA-INV/0027/2018, CSAINV20-nov-0021 to M.L.K.C.).
We thank all of the participants and their families in this study.
We also thank Innovent Pharmaceutical for providing free sintilimab and logistical support for all participants; J.
T.
S.
Wee and T.
S.
Huey for their insightful comments on the trial design; and Y.-X.
Li, X.-Z.
Chen and Y.
Guo for their contributions as members of the independent data monitoring committee.
The trial sponsors did not have roles in data collection, interpretation or manuscript preparation.
These authors contributed equally: Jiawei Lv, Dan-Xue Zheng, Jin-Hui Liang, Ning Zhang, Zu-Lu Ye, Xu-Dong Xu, Melvin.
L.
K.
Chua
These authors jointly supervised this work.
Ji-Bin Li, Guan-Qun Zhou, Jun Ma, Ying Sun
Department of Radiation Oncology, Sun Yat-sen University Cancer Centre, State Key Laboratory of Oncology in South China, Guangdong Key Laboratory of Nasopharyngeal Carcinoma Diagnosis and Therapy, Guangdong Provincial Clinical Research Centre for Cancer, Guangzhou, P.
R.
China
Jiawei Lv
( 吕佳蔚 ), Dan-Xue Zheng
( 郑丹雪 ), Xu-Dong Xu
( 许旭东 ), Wen-Fei Li
( 李文斐 ), Ling-Long Tang
( 唐玲珑 ), Lei Chen
( 陈磊 ), Yan-Ping Mao
( 毛燕萍 ), Rui Guo
( 郭蕊 ), Yu-Pei Chen
( 陈雨沛 ), Li Lin
( 林丽 ), Yuan Zhang
( 张媛 ), Xu Liu
( 刘需 ), Cheng Xu
( 徐骋 ), Zhi-Xuan Li
( 李智轩 ), Ling-Xin Xu
( 徐凌芯 ), Pan-Yang Yang
( 杨潘阳 ), Kun Chen
( 陈昆 ), Guan-Qun Zhou
( 周冠群 ), Jun Ma
( 马骏 ) & Ying Sun
( 孙颖 )
Department of Radiation Oncology, Wuzhou Red Cross Hospital, Wuzhou, P.
R.
China
Jin-Hui Liang
( 梁锦辉 ), Deng Bin
( 邓滨 ), Tian-Sheng Gao
( 高天生 ) & Jian-Ye Yan
( 颜建业 )
Department of Radiation Oncology, First People’s Hospital of Foshan, Foshan, P.
R.
China
Ning Zhang
( 张宁 ) & Lu-Si Chen
( 陈露斯 )
Department of Molecular Diagnostics, Sun Yat-sen University Cancer Center, Guangzhou, P.
R.
China
Zu-Lu Ye
( 叶祖禄 ), Lu-Lu Zhang
( 张露露 ), Zi-Ming Du
( 杜紫明 ) & Zi-Chen Zhang
( 张子辰 )
Duke NUS Medical School and Division of Radiation Oncology, National Cancer Centre, Singapore, Singapore
Melvin.
L.
K.
Chua
( 蔡利强 )
Department of Radiation Oncology, Princess Margaret Cancer Centre, University of Toronto, Toronto, Ontario, Canada
Shao Hui Huang
( 黄少翬 )
Department of Medical Oncology and Clinical Research, Sun Yat-sen University Cancer Center, Guangzhou, P.
R.
China
Hong-Yun Zhao
( 赵洪云 )
Department of Endocrinology, First Affiliated Hospital, Sun Yat-sen University, Guangzhou, P.
R.
China
Shu-Bin Hong
( 洪澍彬 )
Department of Infectious Diseases, Third Affiliated Hospital, Sun Yat-sen University, Guangzhou, P.
R.
China
Yu-Sheng Jie
( 揭育胜 )
Department of Cardiology, Cardiac Prevention and Assessment Center, First Affiliated Hospital, Sun Yat-sen University, Guangzhou, P.
R.
China
Hui-Ling Huang
( 黄慧玲 )
Department of Dermatology, First Affiliated Hospital, Sun Yat-sen University, Guangzhou, P.
R.
China
Xu-Hua Tang
( 唐旭华 )
Department of Pathology, Sun Yat-sen University Cancer Center, Guangzhou, P.R.
China
Jing-Ping Yun
( 云径平 )
Department of Radiology, Sun Yat-sen University Cancer Center, Guangzhou, P.R.
China
Li-Zhi Liu
( 刘立志 ), Li Tian
( 田丽 ) & Hao-Jiang Li
( 黎浩江 )
Department of Clinical Research, Sun Yat-sen University Cancer Centre, Guangzhou, P.
R.
China
Search author on: PubMed Google Scholar
The trial was investigator-initiated and designed by Y.S.
and J.L.
Lead investigators from each centre (Y.S., J.-H.
L.
and N.Z.) contributed to patient recruitment and data acquisition.
J.L., D.-X.Z., Z.-L.Y., J.-B.L., J.M.
and Y.S.
drafted the article and performed the data interpretation.
J.L., D.-X.Z., Z.-L.Y., X.-D.X., M.L.K.C., S.H.H., J.-B.L., G.-Q.Z., J.M.
and Y.S.
revised the manuscript.
G.-Q.Z., L.-L.T., W.-F.L., L-S.C., B.D., T.-S.G., J.-Y.Y., L.C., Y.-P.M., R.G., L.L., Y.-P.C., Y.Z., X.L., Z.-X.L., L.-X.X., P.-Y.Y., K.C., H.-Y.Z., Y.-S.J., H.-L.H.
and X.-H.T.
contributed to clinical management and/or QoL surveys.
Z.-L.Y., Z.-C.Z., Z.-M.D.
and L.-L.Z.
contributed to ctDNA testing.
J.-P.Y., L.-Z.L., L.T.
and H.-J.L.
contributed to the diagnosis and efficacy evaluation of patients.
All authors reviewed, revised and approved the final manuscript.
Correspondence to Ji-Bin Li
( 李济宾 ) , Guan-Qun Zhou
( 周冠群 ) , Jun Ma
( 马骏 ) or Ying Sun
( 孙颖 ) .
M.L.K.C.
reports personal fees for advisory board and education activities and funding support from BeiGene (tislelizumab) for an investigator-initiated trial; personal fees from TopAlliance Biosciences (toripalimab) for advisory board activities; personal fees from Astellas, Pfizer, MSD, AstraZeneca, Varian, Janssen, IQVIA and Telix Pharmaceuticals; nonfinancial support from AstraZeneca; nonfinancial support from Veracyte; and personal fees and grants from Bayer.
M.L.K.C.
consults for ImmunoSCAPE; is a co-inventor on the patent ‘High sensitivity lateral flow immunoassay for detection of analyte in sample’ (10202107837T, Singapore); and serves on the board of directors of Digital Life Line, which owns the licensing agreement of the patent.
The remaining authors declare no competing interests.
Nature thanks the anonymous reviewers for their contribution to the peer review of this work.
Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig.
1 Blood sample collection for ctDNA assessment.
Blood samples for ctDNA assessment were collected at pretreatment (T0), after each cycle of GP-NAC (T1–T3, 21 days after the prior cycle and before the next), and weekly during the adaptive phase (T4–T15) for patients with detectable ctDNA, continuing until clearance was confirmed in two consecutive tests).
At the completion of CCRT, ctDNA assessment was conducted for all enrolled patients.
DDP, cisplatin; RT, radiotherapy.
Extended Data Fig.
2 Outline of the participant selection process in the no-RAT external cohort.
Patients from the EP-SEASON study who met the same eligibility criteria as the EP-STAR trial and received SOC without modifications were included as a pre-planned non-randomized external cohort for efficacy comparison.
Extended Data Fig.
3 Secondary survival outcomes of patients in the intention-to-treat population.
a – c , Kaplan–Meier curves of OS ( a ), DMFS ( b ) and LRFS ( c ) in the RAT group and SOC arm.
Extended Data Fig.
4 Biological correlative analysis of the risk-adaptive interventions across ctDNA-defined risk subgroups.
a , Violin plot showing the signature score for chemotherapy sensitivity across ctDNA-defined risk subgroups (n = 4 samples for low-risk subgroup, n = 6 samples for intermediate-risk subgroup, and n = 5 samples for high-risk subgroup).
b , Venn plot showing the overlap of upregulated genes after GP-NAC across ctDNA-defined risk subgroups.
c , Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment plot of tumour cell upregulated signalling pathways after GP-NAC in low-risk patients.
d , Percentage of innate-like B cells (ILBs) after GP-NAC in low-risk patients (n = 4 pairs).
e , Violin plot showing the signature score for cytotoxic T cells after GP-NAC in low-risk patients (n = 4 pairs).
f , Violin plot showing the signature score for CSC after GP-NAC in intermediate-risk patients (n = 4 samples for low-risk subgroup, n = 6 samples for intermediate-risk subgroup, and n = 5 samples for high-risk subgroup).
g , Violin plot showing the signature score for exhaustion T cells after GP-NAC in high-risk patients (n = 4 samples for low-risk subgroup, n = 6 samples for intermediate-risk subgroup, and n = 5 samples for high-risk subgroup).
The box plot indicates the median (centre), 25th and 75th percentiles (box boundaries), and minimum and maximum (the whiskers) in a , d – g .
Significance was determined by a two-sided Wilcoxon rank-sum test for a , e – g , a two-sided, hypergeometric test for c without correction for multiple comparisons and a two-sided t -test for d .
Extended Data Table 1 Comparison of baseline characteristics of the at-risk and low-risk populations in the EP-STAR trial and the no-RAT external cohort Full size table
Extended Data Table 2 Mean differences of QoL scores in the RAT group versus the SOC arm during the RAT phase Full size table
Extended Data Table 3 Cost-effectiveness analyses of the ctDNA-guided RAT strategy versus the conventional fixed-course, static treatment approach Full size table
Extended Data Table 4 Clinical scenario analysis of cost-effectiveness under an alternative standard, fixed-course, static treatment Full size table
Supplementary Information (download PDF )
This file contains Supplementary Methods, Supplementary Tables 1–26 and Supplementary Figs.
1−4
Reporting Summary (download PDF )
Supplementary Note (download PDF )
Supplementary Data (download ZIP )
Source data for Supplementary Figs.
1, 2 and 4
Peer Review file (download PDF )
Source Data Fig.
3 (download XLSX )
Source Data Fig.
4 (download XLSX )
Source Data Extended Data Fig.
3 (download XLSX )
Source Data Extended Data Fig.
4 (download XLSX )
Springer Nature or its licensor (e.g.
a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
Version of record : 11 March 2026
DOI : https://doi.org/10.1038/s41586-026-10244-w
Related Stories
Source: This article was originally published by Nature News
Read Full Original Article →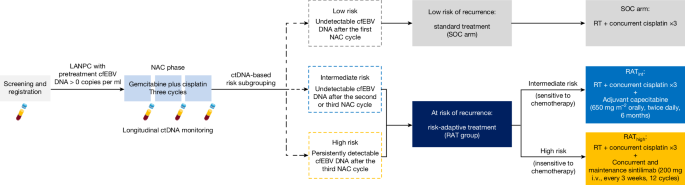



Comments (0)
No comments yet. Be the first to comment!
Leave a Comment