Nature ( 2026 ) Cite this article
Genome-editing technologies that use recombinases to insert kilobase-scale DNA sequences into mammalian genomes canonically require large double-stranded DNA (dsDNA) donors 1 , 2 .
However, dsDNA molecules evoke problematic and toxic innate immune responses, limiting integration efficiencies and generally constraining applicability to ex vivo or immune-deficient contexts.
By harnessing mechanisms of integrative prokaryotic viruses and mobile genetic elements, here we demonstrate that recombinases are compatible with immune evasive circular single-stranded DNA molecules optimally bearing a partial-duplex region that reconstitutes the recombinase recognition sequence.
This approach, which we term integration through nucleus-synthesized template addition of large lengths (INSTALL), is compatible with diverse protein and RNA-guided recombinases for high-fidelity kilobase-scale human genome writing.
INSTALL minimizes innate immune responses in primary human cells and in mice, improving recombinase-mediated integration efficiencies and supporting systemic in vivo non-viral DNA delivery by substantially increasing tolerability and broadening the dosing range compared with lipid nanoparticle-delivered dsDNA molecules.
Together, INSTALL overcomes fundamental challenges for DNA delivery and integration methods by synergizing immune-stealth nucleic acids with recombinases to enable kilobase-scale integration strategies without viral vectors.
This is a preview of subscription content, access via your institution
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
Receive 51 print issues and online access
Prices may be subject to local taxes which are calculated during checkout
Fig.
1: Mechanisms and comparison of the immune-provoking nature of dsDNA and cssDNA molecules.
Fig.
2: Development and characterization of INSTALL.
Fig.
3: INSTALL-mediated DNA integration in diverse human cells and in vivo.
Fig.
4: INSTALL-mediated immune evasion in vitro and in vivo.
Similar content being viewed by others
Efficient non-viral immune cell engineering using circular single-stranded DNA-mediated genomic integration
STRAIGHT-IN: a platform for rapidly generating panels of genetically modified human pluripotent stem cell lines
Systematic discovery of recombinases for efficient integration of large DNA sequences into the human genome
Primary datasets are available in Supplementary Table 6 and RNA-seq and long-read sequencing results are available at the NCBI Sequence Read Archive under accession number PRJNA1296083 .
ClinVar was accessed online ( https://www.ncbi.nlm.nih.gov/clinvar ).
Gaj, T., Sirk, S.
J.
& Barbas, C.
F.
Expanding the scope of site-specific recombinases for genetic and metabolic engineering.
Biotechnol.
Bioeng.
111 , 1–15 (2014).
Article CAS PubMed Google Scholar
Olorunniji, F.
J., Rosser, S.
J.
& Stark, W.
M.
Site-specific recombinases: molecular machines for the genetic revolution.
Biochem.
J.
473 , 673–684 (2016).
Meinke, G., Bohm, A., Hauber, J., Pisabarro, M.
T.
& Buchholz, F.
Cre recombinase and other tyrosine recombinases.
Chem.
Rev.
116 , 12785–12820 (2016).
Jelicic, M.
et al.
Discovery and characterization of novel Cre-type tyrosine site-specific recombinases for advanced genome engineering.
Nucleic Acids Res.
51 , 5285–5297 (2023).
Article CAS PubMed PubMed Central Google Scholar
Smith, M.
C.
M., Brown, W.
R.
A., McEwan, A.
R.
& Rowley, P.
A.
Site-specific recombination by φC31 integrase and other large serine recombinases.
Biochem.
Soc.
Trans.
38 , 388–394 (2010).
Xu, Z.
et al.
Accuracy and efficiency define Bxb1 integrase as the best of fifteen candidate serine recombinases for the integration of DNA into the human genome.
BMC Biotechnol.
13 , 87 (2013).
Article PubMed PubMed Central Google Scholar
Yarnall, M.
T.
N.
et al.
Drag-and-drop genome insertion of large sequences without double-strand DNA cleavage using CRISPR-directed integrases.
Nat.
Biotechnol.
41 , 500–512 (2023).
Durrant, M.
G.
et al.
Systematic discovery of recombinases for efficient integration of large DNA sequences into the human genome.
Nat.
Biotechnol.
41 , 488–499 (2023).
Wu, S.
C.-Y.
et al.
piggyBac is a flexible and highly active transposon as compared to Sleeping Beauty , Tol2 , and Mos1 in mammalian cells.
Proc.
Natl Acad.
Sci.
USA 103 , 15008–15013 (2006).
Article ADS CAS PubMed PubMed Central Google Scholar
Yusa, K., Zhou, L., Li, M.
A., Bradley, A.
& Craig, N.
L.
A hyperactive piggyBac transposase for mammalian applications.
Proc.
Natl Acad.
Sci.
USA 108 , 1531–1536 (2011).
Klompe, S.
E., Vo, P.
L.
H., Halpin-Healy, T.
S.
& Sternberg, S.
H.
Transposon-encoded CRISPR–Cas systems direct RNA-guided DNA integration.
Nature 571 , 219–225 (2019).
Strecker, J.
et al.
RNA-guided DNA insertion with CRISPR-associated transposases.
Science 365 , 48–53 (2019).
Durrant, M.
G.
et al.
Bridge RNAs direct programmable recombination of target and donor DNA.
Nature 630 , 984–993 (2024).
Siddiquee, R., Pong, C.
H., Hall, R.
M.
& Ataide, S.
F.
A programmable seekRNA guides target selection by IS1111 and IS110 type insertion sequences.
Nat.
Commun.
15 , 5235 (2024).
Mukhametzyanova, L.
et al.
Activation of recombinases at specific DNA loci by zinc-finger domain insertions.
Nat.
Biotechnol.
https://doi.org/10.1038/s41587-023-02121-y (2024).
Fauser, F.
et al.
Systematic development of reprogrammed modular integrases enables precise genomic integration of large DNA Sequences.
Preprint at bioRxiv https://doi.org/10.1101/2024.05.09.593242 (2024).
Merrick, C.
A., Zhao, J.
& Rosser, S.
J.
Serine integrases: advancing synthetic biology.
ACS Synth.
Biol.
7 , 299–310 (2018).
Hornung, V.
& Latz, E.
Intracellular DNA recognition.
Nat.
Rev.
Immunol.
10 , 123–130 (2010).
Motwani, M., Pesiridis, S.
& Fitzgerald, K.
A.
DNA sensing by the cGAS–STING pathway in health and disease.
Nat.
Rev.
Genet.
20 , 657–674 (2019).
Carpenter, S.
& O’Neill, L.
A.
J.
From periphery to center stage: 50 years of advancements in innate immunity.
Cell 187 , 2030–2051 (2024).
Sun, L., Wu, J., Du, F., Chen, X.
& Chen, Z.
J.
Cyclic GMP-AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway.
Science 339 , 786–791 (2013).
Article ADS CAS PubMed Google Scholar
Wu, J.
et al.
Cyclic GMP-AMP is an endogenous second messenger in innate immune signaling by cytosolic DNA.
Science 339 , 826–830 (2013).
Neves, C., Escriou, V., Byk, G., Scherman, D.
& Wils, P.
Intracellular fate and nuclear targeting of plasmid DNA.
Cell Biol.
Toxicol.
15 , 193–202 (1999).
Zabner, J., Fasbender, A.
J., Moninger, T., Poellinger, K.
A.
& Welsh, M.
J.
Cellular and molecular barriers to gene transfer by a cationic lipid.
J.
Biol.
Chem.
270 , 18997–19007 (1995).
Wang, D., Tai, P.
W.
L.
& Gao, G.
Adeno-associated virus vector as a platform for gene therapy delivery.
Nat.
Rev.
Drug Discov.
18 , 358–378 (2019).
Hanlon, K.
S.
et al.
High levels of AAV vector integration into CRISPR-induced DNA breaks.
Nat.
Commun.
10 , 4439 (2019).
Article ADS PubMed PubMed Central Google Scholar
Greig, J.
A.
et al.
Integrated vector genomes may contribute to long-term expression in primate liver after AAV administration.
Nat.
Biotechnol.
42 , 1232–1242 (2024).
Simpson, B.
P., Yrigollen, C.
M., Izda, A.
& Davidson, B.
L.
Targeted long-read sequencing captures CRISPR editing and AAV integration outcomes in brain.
Mol.
Ther.
31 , 760–773 (2023).
Ertl, H.
C.
J.
Immunogenicity and toxicity of AAV gene therapy.
Front.
Immunol.
13 , 975803 (2022).
Hinderer, C.
et al.
Severe toxicity in nonhuman primates and piglets following high-dose intravenous administration of an adeno-associated virus vector expressing human SMN.
Hum.
Gene Ther.
29 , 285–298 (2018).
Mays, L.
E.
& Wilson, J.
M.
The complex and evolving story of T cell activation to AAV vector-encoded transgene products.
Mol.
Ther.
19 , 16–27 (2011).
Kotterman, M.
A., Chalberg, T.
W.
& Schaffer, D.
V.
Viral vectors for gene therapy: translational and clinical outlook.
Annu.
Rev.
Biomed.
Eng.
17 , 63–89 (2015).
Moreno, A.
M.
et al.
Immune-orthogonal orthologues of AAV capsids and of Cas9 circumvent the immune response to the administration of gene therapy.
Nat.
Biomed.
Eng.
3 , 806–816 (2019).
Wu, Z., Yang, H.
& Colosi, P.
Effect of genome size on AAV vector packaging.
Mol.
Ther.
18 , 80–86 (2010).
Manno, C.
S.
et al.
Successful transduction of liver in hemophilia by AAV-factor IX and limitations imposed by the host immune response.
Nat.
Med.
12 , 342–347 (2006).
Chirmule, N.
et al.
Humoral immunity to adeno-associated virus type 2 vectors following administration to murine and nonhuman primate muscle.
J.
Virol.
74 , 2420–2425 (2000).
Das, A.
et al.
Epigenetic silencing of recombinant adeno-associated virus genomes by NP220 and the HUSH complex.
J.
Virol.
96 , e02039-21 (2022).
Cabrera, A.
et al.
The sound of silence: transgene silencing in mammalian cell engineering.
Cell Syst.
13 , 950–973 (2022).
Wang, J.-H., Gessler, D.
J., Zhan, W., Gallagher, T.
L.
& Gao, G.
Adeno-associated virus as a delivery vector for gene therapy of human diseases.
Signal Transduct.
Target.
Ther.
9 , 78 (2024).
Decout, A., Katz, J.
D., Venkatraman, S.
& Ablasser, A.
The cGAS–STING pathway as a therapeutic target in inflammatory diseases.
Nat.
Rev.
Immunol.
21 , 548–569 (2021).
Xie, K.
et al.
Efficient non-viral immune cell engineering using circular single-stranded DNA-mediated genomic integration.
Nat.
Biotechnol.
https://doi.org/10.1038/s41587-024-02504-9 (2024).
Cao, X., Tang, L.
& Song, J.
Circular single-stranded DNA: discovery, biological effects, and applications.
ACS Synth.
Biol.
13 , 1038–1058 (2024).
Tang, L.
et al.
Circular single-stranded DNA as switchable vector for gene expression in mammalian cells.
Nat.
Commun.
14 , 6665 (2023).
Kranzusch, P.
J., Lee, A.
S.-Y., Berger, J.
M.
& Doudna, J.
A.
Structure of human cGAS reveals a conserved family of second-messenger enzymes in innate immunity.
Cell Rep.
3 , 1362–1368 (2013).
Kumari, P., Russo, A.
J., Shivcharan, S.
& Rathinam, V.
A.
AIM2 in health and disease: Inflammasome and beyond.
Immunol.
Rev.
297 , 83–95 (2020).
Roberts, T.
L.
et al.
HIN-200 proteins regulate caspase activation in response to foreign cytoplasmic DNA.
Science 323 , 1057–1060 (2009).
Unterholzner, L.
et al.
IFI16 is an innate immune sensor for intracellular DNA.
Nat.
Immunol.
11 , 997–1004 (2010).
Garg, A., Stallings, C.
M.
& Sohn, J.
Filament assembly underpins the double-stranded DNA specificity of AIM2-like receptors.
Nucleic Acids Res.
51 , 2574–2585 (2023).
Krupovic, M.
& Forterre, P.
Single-stranded DNA viruses employ a variety of mechanisms for integration into host genomes.
Ann.
N.
Y.
Acad.
Sci.
1341 , 41–53 (2015).
Roth, T.
L.
et al.
Reprogramming human T cell function and specificity with non-viral genome targeting.
Nature 559 , 405–409 (2018).
Shy, B.
R.
et al.
High-yield genome engineering in primary cells using a hybrid ssDNA repair template and small-molecule cocktails.
Nat.
Biotechnol.
41 , 521–531 (2023).
Iyer, S.
et al.
Efficient homology-directed repair with circular single-stranded DNA donors.
CRISPR J.
https://doi.org/10.1089/crispr.2022.0058 (2022).
Yeh, C.
D., Richardson, C.
D.
& Corn, J.
E.
Advances in genome editing through control of DNA repair pathways.
Nat.
Cell Biol.
21 , 1468–1478 (2019).
Tou, C.
J.
& Kleinstiver, B.
P.
Recent advances in double-strand break-free kilobase-scale genome editing technologies.
Biochemistry https://doi.org/10.1021/acs.biochem.2c00311 (2022).
Shepherd, T.
R., Du, R.
R., Huang, H., Wamhoff, E.-C.
& Bathe, M.
Bioproduction of pure, kilobase-scale single-stranded DNA.
Sci.
Rep.
9 , 6121 (2019).
Lucas, C.
R.
et al.
DNA origami nanostructures elicit dose-dependent immunogenicity and are nontoxic up to high doses in vivo.
Small 18 , 2108063 (2022).
Ranawakage, D.
C.
et al.
Efficient CRISPR-Cas9-mediated knock-in of composite tags in zebrafish using long ssDNA as a donor.
Front.
Cell Dev.
Biol.
8 , 598634 (2021).
Lin-Shiao, E.
et al.
CRISPR–Cas9-mediated nuclear transport and genomic integration of nanostructured genes in human primary cells.
Nucleic Acids Res.
50 , 1256–1268 (2022).
Andreeva, L.
et al.
cGAS senses long and HMGB/TFAM-bound U-turn DNA by forming protein–DNA ladders.
Nature 549 , 394–398 (2017).
Luecke, S.
et al.
cGAS is activated by DNA in a length-dependent manner.
EMBO Rep.
18 , 1707–1715 (2017).
Zhou, W.
et al.
Structure of the human cGAS–DNA complex reveals enhanced control of immune surveillance.
Cell 174 , 300–311 (2018).
Reus, J.
B., Trivino-Soto, G.
S., Wu, L.
I., Kokott, K.
& Lim, E.
S.
SV40 large T antigen is not responsible for the loss of STING in 293T cells but can inhibit cGAS-STING interferon induction.
Viruses 12 , 137 (2020).
Bonnamy, M., Blanc, S.
& Michalakis, Y.
Replication mechanisms of circular ssDNA plant viruses and their potential implication in viral gene expression regulation.
mBio 14 , e01692-23 (2023).
Geider, K.
& Kornberg, A.
Conversion of the M13 viral single strand to the double-stranded replicative forms by purified proteins.
J.
Biol.
Chem.
249 , 3999–4005 (1974).
Fanton, A.
et al.
Site-specific DNA insertion into the human genome with engineered recombinases.
Nat.
Biotechnol.
https://doi.org/10.1038/s41587-025-02895-3 (2025).
Tobiason, D.
M., Lenich, A.
G.
& Glasgow, A.
C.
Multiple DNA binding activities of the novel site-specific recombinase, Piv, from Moraxella lacunata .
J.
Biol.
Chem.
274 , 9698–9706 (1999).
Marrs, C.
F., Rozsa, F.
W., Hackel, M., Stevens, S.
P.
& Glasgow, A.
C.
Identification, cloning, and sequencing of piv, a new gene involved in inverting the pilin genes of Moraxella lacunata .
J.
Bacteriol.
172 , 4370–4377 (1990).
Lau, L., Gray, E.
E., Brunette, R.
L.
& Stetson, D.
B.
DNA tumor virus oncogenes antagonize the cGAS-STING DNA-sensing pathway.
Science 350 , 568–571 (2015).
Takaki, T., Millar, R., Hiley, C.
T.
& Boulton, S.
J.
Micronuclei induced by radiation, replication stress, or chromosome segregation errors do not activate cGAS-STING.
Mol.
Cell 84 , 2203–2213 (2024).
Al Khatib, I.
et al.
Activation of the cGAS-STING innate immune response in cells with deficient mitochondrial topoisomerase TOP1MT.
Hum.
Mol.
Genet.
32 , 2422–2440 (2023).
Chen, G.-Y.
et al.
Defective antiviral responses of induced pluripotent stem cells to baculoviral vector transduction.
J.
Virol.
86 , 8041–8049 (2012).
Anzalone, A.
V.
et al.
Programmable deletion, replacement, integration and inversion of large DNA sequences with twin prime editing.
Nat.
Biotechnol.
40 , 731–740 (2022).
Anzalone, A.
V.
et al.
Search-and-replace genome editing without double-strand breaks or donor DNA.
Nature 576 , 149–157 (2019).
Ferreira Da Silva, J.
et al.
Click editing enables programmable genome writing using DNA polymerases and HUH endonucleases.
Nat.
Biotechnol.
https://doi.org/10.1038/s41587-024-02324-x (2024).
Liu, B.
et al.
Targeted genome editing with a DNA-dependent DNA polymerase and exogenous DNA-containing templates.
Nat.
Biotechnol.
https://doi.org/10.1038/s41587-023-01947-w (2023).
Pandey, S.
et al.
Efficient site-specific integration of large genes in mammalian cells via continuously evolved recombinases and prime editing.
Nat.
Biomed.
Eng.
https://doi.org/10.1038/s41551-024-01227-1 (2024).
Karasu, M.
E.
et al.
Removal of TREX1 activity enhances CRISPR–Cas9-mediated homologous recombination.
Nat.
Biotechnol.
https://doi.org/10.1038/s41587-024-02356-3 (2024).
Yan, J.
et al.
Improving prime editing with an endogenous small RNA-binding protein.
Nature 628 , 639–647 (2024).
Hazelbaker, D.
Z.
et al.
Large serine integrase off-target discovery and validation for therapeutic genome editing.
Preprint at bioRxiv https://doi.org/10.1101/2024.08.23.609471 (2024).
Giannoukos, G.
et al.
UDiTaS TM , a genome editing detection method for indels and genome rearrangements.
BMC Genom.
19 , 212 (2018).
Adams, D.
et al.
Patisiran, an RNAi therapeutic, for hereditary transthyretin amyloidosis.
N.
Engl.
J.
Med.
379 , 11–21 (2018).
Low, B.
E., Hosur, V., Lesbirel, S.
& Wiles, M.
V.
Efficient targeted transgenesis of large donor DNA into multiple mouse genetic backgrounds using bacteriophage Bxb1 integrase.
Sci.
Rep.
12 , 5424 (2022).
Rose, J.
et al.
Engineered Bxb1 variants improve integrase activity and fidelity.
Preprint at bioRxiv https://doi.org/10.1101/2024.10.21.619419 (2024).
Dong, M.
& Fitzgerald, K.
A.
DNA-sensing pathways in health, autoinflammatory and autoimmune diseases.
Nat.
Immunol.
25 , 2001–2014 (2024).
Patel, M.
N.
et al.
Safer non-viral DNA delivery using lipid nanoparticles loaded with endogenous anti-inflammatory lipids.
Nat.
Biotechnol.
https://doi.org/10.1038/s41587-025-02556-5 (2025).
Roberts, T.
C., Langer, R.
& Wood, M.
J.
A.
Advances in oligonucleotide drug delivery.
Nat.
Rev.
Drug Discov.
19 , 673–694 (2020).
Yin, H.
et al.
Structure-guided chemical modification of guide RNA enables potent non-viral in vivo genome editing.
Nat.
Biotechnol.
35 , 1179–1187 (2017).
Finn, J.
D.
et al.
A single administration of CRISPR/Cas9 lipid nanoparticles achieves robust and persistent in vivo genome editing.
Cell Rep.
22 , 2227–2235 (2018).
Hew, B.
E.
et al.
Directed evolution of hyperactive integrases for site specific insertion of transgenes.
Nucleic Acids Res.
52 , e64 (2024).
Huhtinen, O., Prince, S., Lamminmäki, U., Salbo, R.
& Kulmala, A.
Increased stable integration efficiency in CHO cells through enhanced nuclear localization of Bxb1 serine integrase.
BMC Biotechnol 24 , 44 (2024).
Karikó, K., Buckstein, M., Ni, H.
& Weissman, D.
Suppression of RNA recognition by toll-like receptors: the impact of nucleoside modification and the evolutionary origin of RNA.
Immunity 23 , 165–175 (2005).
Cha, T.-G.
et al.
Genetic control of aerogel and nanofoam properties, applied to Ni–MnO cathode design.
Adv.
Funct.
Mater.
31 , 2010867 (2021).
Wrenbeck, E.
E.
et al.
Plasmid-based one-pot saturation mutagenesis.
Nat.
Methods 13 , 928–930 (2016).
Kim, M.
J.
& Ahituv, N.
in Pharmacogenomics: Methods and Protocols (eds Innocenti, F.
& van Schaik, R.
H.
N.) 279–289 (Humana Press, 2013).
Rohland, N.
& Reich, D.
Cost-effective, high-throughput DNA sequencing libraries for multiplexed target capture.
Genome Res.
22 , 939–946 (2012).
Kleinstiver, B.
P.
et al.
Engineered CRISPR–Cas12a variants with increased activities and improved targeting ranges for gene, epigenetic and base editing.
Nat.
Biotechnol.
37 , 276–282 (2019).
Chen, P.
J.
et al.
Enhanced prime editing systems by manipulating cellular determinants of editing outcomes.
Cell 184 , 5635–5652.e29 (2021).
Ge, S.
X., Jung, D.
& Yao, R.
ShinyGO: a graphical gene-set enrichment tool for animals and plants.
Bioinformatics 36 , 2628–2629 (2020).
Escudero, J.
A.
et al.
Unmasking the ancestral activity of integron integrases reveals a smooth evolutionary transition during functional innovation.
Nat.
Commun.
7 , 10937 (2016).
Francia, M.
V., Zabala, J.
C., De La Cruz, F.
& García Lobo, J.
M.
The IntI1 integron integrase preferentially binds single-stranded DNA of the attC site.
J.
Bacteriol.
181 , 6844–6849 (1999).
Grabundzija, I., Hickman, A.
B.
& Dyda, F.
Helraiser intermediates provide insight into the mechanism of eukaryotic replicative transposition.
Nat.
Commun.
9 , 1278 (2018).
Del Pilar Garcillán-Barcia, M., Bernales, I., Mendiola, M.
V.
& De La Cruz, F.
Single-stranded DNA intermediates in IS 91 rolling-circle transposition.
Mol.
Microbiol.
39 , 494–502 (2001).
Barabas, O.
et al.
Mechanism of IS200/IS605 family DNA transposases: activation and transposon-directed target site selection.
Cell 132 , 208–220 (2008).
Xie, K.
et al.
Efficient non-viral immune cell engineering using circular single-stranded DNA-mediated genomic integration.
Nat.
Biotechnol.
43 , 1821–1832 (2025).
Hew, B.
E.
et al.
Directed evolution of hyperactive integrases for site specific insertion of transgenes.
Nucleic Acids Res.
https://doi.org/10.1093/nar/gkae534 (2024).
Wu, J., Corbett, A.
H.
& Berland, K.
M.
The intracellular mobility of nuclear import receptors and NLS cargoes.
Biophys.
J.
96 , 3840–3849 (2009).
Suzuki, K.
et al.
In vivo genome editing via CRISPR/Cas9 mediated homology-independent targeted integration.
Nature 540 , 144–149 (2016).
Witte, I.
P.
et al.
Programmable gene insertion in human cells with a laboratory-evolved CRISPR-associated transposase.
Science 388 , eadt5199 (2025).
We thank D.
Wagner and I.
Kassing for sharing protocols related to culturing and activating primary human T cells; M.
Nahrendorf, A.
Paccalet and M.
Hulsmans for advice related to THP-1 cells and cytokine analyses; E.
King for assistance with molecular cloning; M.
Wiles, S.
Murray and B.
Low from Jackson Laboratory for advice on mouse models; S.
Levesque and D.
Bauer for advice on La-compatible pegRNA designs; J.
Owens for discussions on engineered Bxb1 enzymes; R.
Xu, Z.
Zhu and S.
Zhao from Quintara Biosciences for DNA synthesis, plasmid cloning and DNA/RNA-seq services; Azenta for assisting with and expediting Olink analyses.
We acknowledge funding from a National Science Foundation Graduate Research Fellowship (2020295403; C.J.T.), an EMBO Long Term Fellowship (ALTF 750-2022; to J.F.d.S.), a Friedreich’s Ataxia Research Alliance (FARA) and FARA Australia Fellowship (D.R.-R.), the Kayden–Lambert MGH Research Scholar Award 2023–2028 (B.P.K) and National Institutes of Health grants and DP2CA281401 (B.P.K.), P01HL142494 (B.P.K.), and R01NS125353 (P.L.M.
and B.P.K.).
These authors contributed equally: Connor J.
Tou, Keqiang Xie
Center for Genomic Medicine, Massachusetts General Hospital, Boston, MA, USA
Connor J.
Tou, Joana Ferreira da Silva, Pazhanichamy Kalailingam, David Rufino-Ramos, Madeline L.
Eller, Patricia L.
Musolino & Benjamin P.
Kleinstiver
Department of Pathology, Massachusetts General Hospital, Boston, MA, USA
Connor J.
Tou, Joana Ferreira da Silva, David Rufino-Ramos, Madeline L.
Eller & Benjamin P.
Kleinstiver
Biological Engineering Program, Massachusetts Institute of Technology, Cambridge, MA, USA
Full Circles Therapeutics, Cambridge, MA, USA
Keqiang Xie, Jakob Starzyk, Ishita Majumdar, Jiao Wang, Danna Lee, Richard Shan & Hao Wu
Department of Pathology, Harvard Medical School, Boston, MA, USA
Joana Ferreira da Silva, David Rufino-Ramos & Benjamin P.
Kleinstiver
Department of Neurology, Massachusetts General Hospital, Boston, MA, USA
Pazhanichamy Kalailingam & Patricia L.
Musolino
Department of Neurology, Harvard Medical School, Boston, MA, USA
Institute for Medical Engineering and Science, Massachusetts Institute of Technology, Cambridge, MA, USA
Eliz Amar-Lewis & Natalie Artzi
Department of Medicine, Division of Engineering in Medicine, Brigham and Women’s Hospital, Harvard Medical School, Boston, MA, USA
Eliz Amar-Lewis, William Sawyer & Natalie Artzi
Wyss Institute for Biologically Inspired Engineering, Harvard University, Boston, MA, USA
Department of Bioengineering, Northeastern University, Boston, MA, USA
Wisconsin Institute for Immune and Cell Therapy, Madison, WI, USA
Stellate DNA, Cambridge, MA, USA
Quintara Bioscience, Cambridge, MA, USA
Search author on: PubMed Google Scholar
C.J.T.
and K.X.
conceived the concept with input from R.S., H.W.
and B.P.K.; C.J.T., K.X., J.F.d.S., J.S., I.M., M.L.E.
and J.W.
designed and performed experiments.
D.R.-R.
generated the attB and attP cell lines.
D.L.
generated cssDNA.
S.Y.
performed experiments in CD-1 mice with guidance from J.L.; E.A.-L.
and W.S.
formulated the LNPs for experiments in attP mice with guidance from N.A.; P.K.
performed experiments in attP mice with guidance from P.L.M.; R.J.M.
and G.A.D.
produced Bxb1 mRNA for experiments in attP mice.
H.W.
and B.P.K.
oversaw the study.
C.J.T.
and B.P.K.
wrote the manuscript with input from all of the other authors.
Correspondence to Connor J.
Tou , Hao Wu or Benjamin P.
Kleinstiver .
C.J.T.
and B.P.K.
are listed as inventors on a patent application filed by Mass General Brigham (MGB) that describes INSTALL.
C.J.T., J.F.d.S.
and B.P.K.
are listed as inventors on additional patents or patent applications filed by MGB that describe genome engineering technologies related to the current study.
K.X., R.S.
and H.W.
are listed as inventors on patent applications filed by Full Circles Therapeutics that describe the production and use of cssDNA.
K.X., J.S., I.M., J.W., D.L., R.S.
and H.W.
are current or former employees of Full Circles Therapeutics.
P.L.M.
holds scientific research agreements with Angea Biotherapeutics and Minoryx and has provided consultation for Capsida, Astellas, Ionis, Biogen, Vertex, Bluebird, Atlas Ventures and Inozyme.
P.L.M.
reports grants to Massachusetts General Hospital from the NIH for research on ALD and genome editing on genetic vasculopathies and extracellular matrix disorders.
B.P.K.
is a consultant for Novartis Venture Fund, Generation Bio and Jumble Therapeutics, and is on the scientific advisory boards of Life Edit Therapeutics and Prime Medicine.
B.P.K.
has a financial interest in Prime Medicine, a company developing therapeutic CRISPR–Cas technologies for gene editing.
B.P.K.’s interests were reviewed and are managed by MGH and MGB in accordance with their conflict-of-interest policies.
The other authors declare no competing interests.
Nature thanks Jesse Owens and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig.
1 Methods of cssDNA propagation, synthetic cssDNA production, and INSTALL-mediated integration in human cells.
( a ) Schematics of integrative lifecycles of viruses with cssDNA genomes that leverage either virally- or host-encoded recombinases for genomic integration before or after second-strand synthesis of their genome by a host DNA polymerase (DNAP) 49 .
( b ) Schematic of integron mobilization and integration into a host bacteria, which uses a cssDNA intermediate with an intramolecular dsDNA region for integration, with second strand synthesis of the genome occurring following integration 99 , 100 .
( c ) Schematic of proposed propagation mechanisms of Helitrons and IS91 (top) elements 101 , 102 or IS200/605 elements 103 (bottom), which utilizes ssDNA transposon intermediates in their integration process.
( d ) Schematic of M13 phage based cssDNA production workflow 55 , 104 ( e ) Representative agarose gel image of cssDNA resulting from the M13 phage production shown in panel d (cssDNA shown as n = 2 replicates; molecular weight marker shown in kilobases).
( f ) Schematic of an optimized enzymatic “nick and digest” method to produce cssDNA from dsDNA starting points, adapted from a previously described method 93 .
( g ) Gel analysis of DNA before (dsDNA plasmid) and after (resulting cssDNA) the “nick and digest” production method shown in panel f (single representative replicate).
( h ) Integration efficiencies when co-delivering bpNLS-Bxb1 mRNA and dsDNA, cssDNA, or oDNA donors into attB HEK 293T cells, measured by droplet digital PCR (ddPCR).
The cssDNA molecules were produced by phagemid production.
( i ) Integration efficiencies when co-delivering bpNLS-Bxb1 mRNA and dsDNA, cssDNA, or oDNA donors into attB HEK 293T cells where either the mRNA and DNA were co-encapsulated in TransIT-mRNA (mixed) or where mRNA was separately encapsulated in TransIT-mRNA and DNA was encapsulated in TransIT-X2 (separate).
Integration efficiencies measured by ddPCR.
For panels h and i , mean, standard deviation (s.d.), and individual datapoints shown for n = 3 independent biological replicates.
Extended Data Fig.
2 Additional biochemical and cellular characterization of INSTALL.
( a ) Schematic of biochemical reactions comprised of different DNA donors and Bxb1 protein derived from human HEK 293T cell lysates.
( b ) Representative PCR products to detect integration from biochemical reactions consisting of Bxb1 protein from HEK 293T cell lysates along with target and donor DNA molecules (single representative replicate); image from a QIAxcel DNA Fast Analysis Kit (Qiagen).
Reactions were incubated for 2 h.
( c ) Integration efficiencies for plasmid targeting integration experiments with dsDNA or cssDNA (INSTALL-1) donors.
( d ) Prior studies 8 identified a conservative Pa01 attP sequence, for which we minimized by testing truncated attP versions.
Integration efficiencies for plasmid expressed Pa01 in attB HEK 293T cells with various lengths of attP sequences on dsDNA donor plasmids, measured by ddPCR.
( e ) Schematic of creating attB and attP landing pad cell lines with attachment sites for various recombinases 8 , via lentiviral transduction with low multiplicity of infection in various human cell types.
( f ) Quantification of lentiviral copies in attB and attP HEK 293T cell lines, measured by ddPCR.
( g ) Junction PCR products to detect full insertions from sequential plasmid delivery experiments with dsDNA, INSTALL-1, or INSTALL-2 (PCR products shown as n = 2 replicates; molecular weight marker shown in kilobases and a 2.2 kb band is indicated).
( h ) Ratio of genome captured DNA copies per genome copy, measured by ddPCR.
HEK 293T cells were transfected with dsDNA or cssDNA and passaged for 4 weeks before quantification.
For panels c, d, f , and h , integration efficiencies measured by ddPCR, mean, s.d., and individual datapoints shown for n = 3 independent biological replicates.
Extended Data Fig.
3 Optimization of nocodazole treatment to arrest HEK 293T cells in the G2/M phase of the cell cycle.
( a ) Simplified schematic of cell cycle phases.
Nocodazole treatment arrests cells in the G2/M phase, which is highlighted.
( b ) Flow cytometry plots of cells treated with various doses of nocodazole and stained with propidium iodide (PI) stain.
For subsequent experiments using nocodazole treatment, a dose of 35 ng/mL was used.
Extended Data Fig.
4 INSTALL with PiggyBac transposase in K562 cells, T-cells, or iPSCs.
( a ) HDR efficiencies measured by ddPCR to place Bxb1 attP site at RAB11A for one-pot integration experiments in primary human T-cells.
( b ) Schematic of INSTALL-1 and INSTALL-2e implementation with PiggyBac transposase.
( c ) GFP+ cells measured by flow cytometry from experiments in K562 cells without or with PiggyBac transposase mRNA and a cssDNA donor containing PiggyBac ITRs and an EF1α-GFP cargo.
( d ) Schematic of dual ePIP approach to partially duplex left and right PiggyBac transposase ITRs and the TTAA motif required for transposon excision.
( e ) GFP+ cells measured by flow cytometry from experiments in immune competent primary human T-cells nucleofected with dsDNA, INSTALL-1, INSTALL-2, or INSTALL-2e.
( f ) Representative FACS plots from PiggyBac experiments in primary human T-cells, as shown in panel e .
( g ) Representative FACS plots from one-pot Bxb1 integration experiments in immune attenuated iPSCs 71 .
Mean, s.d., and individual datapoints shown for n = 5 independent biological replicates ( panel a) or n = 3 independent biological replicates ( panels b and e ).
Extended Data Fig.
5 Bxb1 recombinase engineering and compatibility of INSTALL with prime and click editing.
( a ) Integration efficiencies for dsDNA, cssDNA, or oDNA donors via PASSIGE experiments in immune-deficient HEK 293T cells and with sequential delivery of recombinase plasmid approximately 36 h prior to the donor.
( b ) Schematic of engineered Bxb1 constructs with different NLS configurations, activity-enhancing mutations 76 , 105 , and stabilon or exin21 tags 106 , 107 .
( c-e ) Recombination efficiencies (PE attP site conversion) with Bxb1 variants (shown in panel b ) at high dose (150 ng of donor and 80 ng recombinase expression plasmid; panel c ), moderate dose (20 ng of donor and 30 ng recombinase; panel d ), or low dose (1 ng of donor and 2 ng recombinase; panel e ) of plasmids for PASSIGE experiments in HEK 293T cells (PEmax, pegRNA, Bxb1 variant, and donor plasmids delivered to cells simultaneously).
( f,g ) attP site writing efficiency by twin prime editing (twinPE) ( panel f ) and cargo integration efficiencies ( panel g) for PASSIGE experiments with dsDNA, INSTALL-1, INSTALL-2, or INSTALL-2e donors in primary human T-cells.
( h ) Integration efficiencies from PASSIGE experiments using St.eeBxb1 with a 2 kb or 6 kb oDNA (INSTALL-2e) in HEK 293T cells.
( i ) Long-read sequencing coverage of the 6 kb insertion analysed in panel h .
( j ) Efficiency of Bxb1 attP site writing at the AAVS1 locus using PE7 mRNA, dual synthetic pegRNAs, and different DNA donors in primary human hepatocytes (PHHs), quantified by NGS.
( k,l ) Absolute integration efficiencies and percentage of attP sites recombined ( panels k and l , respectively) in PASSIGE experiments in PHHs when using dsDNA, INSTALL-1, -2, or -2e donors.
Recombinase mRNA and DNA donors were transfected into PHHs 48 h after PE7 components were first transfected, then cells were harvested 3 days later.
( m ) Integration efficiency at on-target or off-target sites for Bxb1 or eeBxb1 when attP is pre-installed in the genome, with off-target events analysed at sites previously nominated by UDiTaS 76 , 80 or HIDE-seq 79 .
( n ) Number of reads that map to on- or off-target sites from PASSIGE experiments in primary human T-cells, assessed via genome-wide integration analysis using an adapted UDiTaS-style assay.
( o ) Comparison of total reads that map to the human genome from our current study or two other recent manuscript that use recombinases or transposases for targeted DNA integration 76 , 108 .
( p ) Proportion of total UDiTaS experiment sequencing reads that map to the DNA donor or to other sources (e.g.
human gDNA, DNA donor with potential sequencing error, etc.).
( q ) Schematic of INSTALL-2 approach to insert a splice acceptor, exons 2-11 of ABCD1 fused to a NanoLuc, and polyA sequence into a HEK 293T cell line Bxb1 attB site in intron 1 of ABCD1 , produced by click editing.
Correct integration leads to expression of the cargo expressed from the endogenous ABCD1 promoter, which can be quantified by ADLP-nLuc protein luminescence.
( r ) Quantification of luminescence from experiments seeking to integrate a INSTALL-1 or -2 donors encoding exons 2-11 of ABCD1 fused to a NanoLuc into the Bxb1 attB site installed in intron 1 of ABCD1 gene in HEK 293T cells.
For panels a , c-h , and m integration efficiencies measured by ddPCR; mean, s.d., and individual datapoints shown for n = 3 independent biological replicates ( panels a , c-e , h , and m ) or n = 4 independent biological replicates ( panels f and g) .
Extended Data Fig.
6 Further characterization of INSTALL-mediated in vivo integration.
( a ) Schematic of LNP components and formation to encapsulate mRNA, dsDNA, or oDNA.
( b-d ) Characterization of LNP properties, including hydrodynamic size (nm) and polydispersity index (PDI) ( panel b ), Zeta potential (mV) ( panel c ), and encapsulation efficiency by the RiboGreen assay ( panel d ).
( e ) Schematic of P3 reto-orbital injections of Bxb1 attP mice 82 with lipid nanoparticles (LNPs) encapsulating mRNA or DNA (left panel) and survival curves for various dsDNA or oDNA doses that also contained a 1 mg/kg bpNLS-Bxb1 mRNA dose (right panel).
n = 5 mice for the 0.35 mg/kg dsDNA dose; n = 3 mice for the 0.5 mg/kg dsDNA dose; and n = 6 mice for the INSTALL-2e condition.
( f ) Images of P3 attP mice dosed with LNPs containing 1.5 mg/kg St.eeBxb1 mRNA and either LNPs containing 0.7 mg/kg dsDNA or 1.4 mg/kg oDNA.
n = 3 mice for all conditions.
( g ) Long-read (full length insertion) and Sanger sequencing (attR and attL junctions) from in vivo integration experiments in neonatal mice when using INSTALL-2e.
( h ) Schematic of adult attP mouse (8-week) tail-vein injections of Bxb1 attP mice 82 with LNPs encapsulating mRNA or DNA (left panel) and survival curves for various dsDNA or oDNA doses that also contained a 1.5 mg/kg St.reBxb1 mRNA dose (right panel).
( i ) Weight change over time of adult attP mice dosed as described in panel h .
( j ) Integration efficiencies measured by ddPCR from in vivo experiments in 8-week old adult attP mice following injection of LNPs containing either dsDNA or INSTALL-2e.
( k ) Sanger sequencing of the attR junction from adult attP mouse experiments when using INSTALL-2e.
For panels b-d , mean, s.d., and individual datapoints are shown.
For panels h-j , mean, s.d., and individual datapoints shown for n = 3 mice.
Extended Data Fig.
7 Analysis of transcriptional perturbations in primary human cells.
( a ) Representative image of THP-1 cell differentiation into macrophages upon addition of 100 ng/µL PMA into the cell culture media.
( b ) Nucleofection efficiency as measured by flow cytometry of GFP+ cells for immune competent primary human T-cells and THP-1 derived macrophages.
Mean, s.d., and individual datapoints shown for n = 2 independent biological replicates.
( c ) Chord plot analysis of the top 16 differentially regulated genes and their associated GO biological process category from primary human T-cells treated with Bxb1 mRNA and dsDNA.
( d, e ) Chord plot analysis of the top 16 differentially regulated genes and their associated GO molecular process category from primary human T-cells ( panel d ) or THP-1 derived macrophages ( panel e ) treated with Bxb1 mRNA and dsDNA.
( f ) Relative IL-6 mRNA in immune competent primary human keratinocytes 6 h after transfection of different doses of equal masses of a CpG-minimized dsDNA, cssDNA, or oDNA donors with Lipofectamine 2000, as measured by RT-qPCR.
For panels b and f , mean and individual datapoints shown for n = 2 independent biological replicates.
Extended Data Fig.
8 Analysis of INSTALL evasion of innate immune activation in liver and blood.
( a ) Images of liver sections stained for CD68+ cells from neonatal (top two rows) or adult (bottom row) mice; P3 neonatal mice were injected retro-orbitally with 1.5 mg/kg St.eeBxb1 mRNA encapsulated in LNPs and either 0.7 or 1.5 mg/kg dsDNA or oDNA encapsulated in LNPs; adult mice were tail vein injected with 1.5 mg/kg St.eeBxb1 mRNA encapsulated in LNPs and 1 mg/kg dsDNA or oDNA encapsulated in LNPs for adult mice.
( b ) Quantification of CD68+ cells from stained images shown in panel a , measured by ImageJ.
( c ) Monocyte percentage and platelet counts from adult attP mice treated with LNPs containing 1.5 mg/kg St.reBxb1 mRNA and LNPs containing 1 mg/kg dsDNA or oDNA.
For panel b and c , mean, s.d., and individual datapoints shown for n = 4 neonatal mice ( panel b ) and n = 3 adult mice ( panel b and c ).
Extended Data Fig.
9 Additional immune marker analysis from in vivo experiments.
( a ) Immune marker levels in neonatal attP mouse plasma after P3 retro-orbital injections of LNPs encapsulating 1.5 mg/kg St.eeBxb1 mRNA and either 0.7 mg/kg dsDNA or oDNA, measured by multiplexed mouse immune marker panel (Olink).
( b ) Schematic of INSTALL-2e immune evasion of TLR9 in the endosome via minimization of CpG motifs on the oDNA donor.
( c ) CpG dinucleotide content in the original oDNA utilized in this study and in a CpG-minimized oDNA.
( d ) Additional immune marker levels in adult attP mouse plasma after treatment with LNPs encapsulating 1.5 mg/kg St.eeBxb1 mRNA and either 1 mg/kg dsDNA or oDNA, measured by a multiplexed FACS-based immune marker panel (BioLegend).
( e ) Summary violin plot of the fold decrease in immune markers when using a CpG-minimized oDNA.
Mean and individual datapoints shown for n = 4 mice for PBS, mRNA, and INSTALL conditions and n = 3 mice for dsDNA condition for panel a and n = 3 mice for all conditions for panel d .
Extended Data Fig.
10 Potential of INSTALL for mutation-agnostic genome editing.
( a ) Histogram of 672 genetic diseases that each contain over 100 known pathogenic variants, identified in the ClinVar database.
( b ) Schematic of one potential use of INSTALL in enabling mutation-agnostic genome editing approaches.
Supplementary Information (download PDF )
Supplementary Figures, Supplementary Notes and Supplementary References.
Reporting Summary (download PDF )
Supplementary Tables 1–6 (download XLSX )
Supplementary Table 1: the plasmids used in this study.
Supplementary Table 2: sequences of PIPs, sgRNAs and pegRNAs.
Supplementary Table 3: ddPCR primers and probes.
Supplementary Table 4: RNA-seq gene expression analysis primary datasets.
Supplementary Table 5: RNA-seq GO enrichment primary datasets.
Supplementary Table 6: primary datasets
Uncropped gel images (download PDF )
Peer Review file (download PDF )
Supplementary Video 1 (download MP4 )
The responses of neonatal attP mice to dsDNA, INSTALL-2e and PBS.
P3 attP mice were retro-orbitally injected with PBS or LNPs containing 1.5 mg per kg St.eeBxb1 mRNA and 1.5 mg per kg dsDNA or oDNA.
Videos were acquired around 30 min after injection.
I2e, INSTALL-2e, including an oDNA donor with annealed ePIP.
Supplementary Video 2 (download MP4 )
The responses of adult attP mice to dsDNA and INSTALL-2e.
8-week-old adult attP mice were tail-vein injected with LNPs containing 1.5 mg per kg St.eeBxb1 mRNA and 1 mg per kg dsDNA or oDNA.
Videos were acquired around 1 h after injection.
I2e, INSTALL-2e.
Supplementary Video 3 (download MP4 )
The responses of adult attP mice to PBS.
8-week-old adult attP mice were injected through tail vein injection with PBS.
Videos were acquired around 1 h after injection
Springer Nature or its licensor (e.g.
a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
Tou, C.J., Xie, K., Ferreira da Silva, J.
et al.
Immune evasive DNA donors and recombinases license kilobase-scale writing.
Nature (2026).
https://doi.org/10.1038/s41586-026-10241-z
Version of record : 11 March 2026
DOI : https://doi.org/10.1038/s41586-026-10241-z
Related Stories
Source: This article was originally published by Nature News
Read Full Original Article →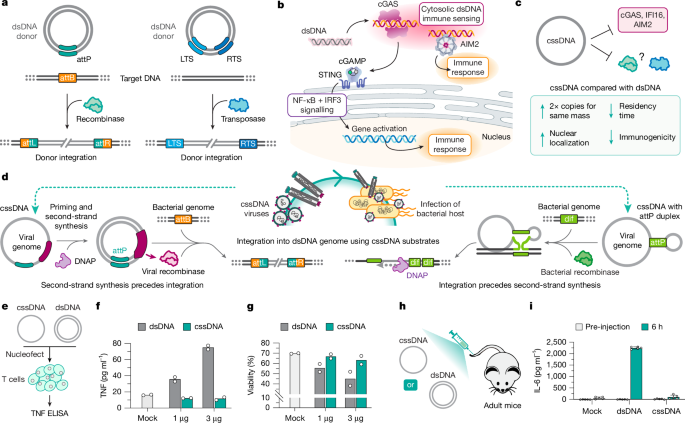



Comments (0)
No comments yet. Be the first to comment!
Leave a Comment